Content from Before we start
Last updated on 2023-06-05 | Edit this page
Estimated time: 30 minutes
Overview
Questions
- What is Python and why should I learn it?
Objectives
- Present motivations for using Python.
- Organize files and directories for a set of analyses as a Python project, and understand the purpose of the working directory.
- How to work with Jupyter Notebook and Spyder.
- Know where to find help.
- Demonstrate how to provide sufficient information for troubleshooting with the Python user community.
What is Python?
Python is a general purpose programming language that supports rapid development of data analytics applications. The word “Python” is used to refer to both, the programming language and the tool that executes the scripts written in Python language.
Its main advantages are:
- Free
- Open-source
- Available on all major platforms (macOS, Linux, Windows)
- Supported by Python Software Foundation
- Supports multiple programming paradigms
- Has large community
- Rich ecosystem of third-party packages
So, why do you need Python for data analysis?
Easy to learn: Python is easier to learn than other programming languages. This is important because lower barriers mean it is easier for new members of the community to get up to speed.
-
Reproducibility: Reproducibility is the ability to obtain the same results using the same dataset(s) and analysis.
Data analysis written as a Python script can be reproduced on any platform. Moreover, if you collect more or correct existing data, you can quickly re-run your analysis!
An increasing number of journals and funding agencies expect analyses to be reproducible, so knowing Python will give you an edge with these requirements.
-
Versatility: Python is a versatile language that integrates with many existing applications to enable something completely amazing. For example, one can use Python to generate manuscripts, so that if you need to update your data, analysis procedure, or change something else, you can quickly regenerate all the figures and your manuscript will be updated automatically.
Python can read text files, connect to databases, and many other data formats, on your computer or on the web.
Interdisciplinary and extensible: Python provides a framework that allows anyone to combine approaches from different research (but not only) disciplines to best suit your analysis needs.
Python has a large and welcoming community: Thousands of people use Python daily. Many of them are willing to help you through mailing lists and websites, such as Stack Overflow and Anaconda community portal.
Free and Open-Source Software (FOSS)… and Cross-Platform: We know we have already said that but it is worth repeating.
Knowing your way around Anaconda
Anaconda distribution of Python includes a lot of its popular packages, such as the IPython console, Jupyter Notebook, and Spyder IDE. Have a quick look around the Anaconda Navigator. You can launch programs from the Navigator or use the command line.
The Jupyter Notebook is an open-source web application that allows you to create and share documents that allow one to create documents that combine code, graphs, and narrative text. Spyder is an Integrated Development Environment that allows one to write Python scripts and interact with the Python software from within a single interface.
Anaconda comes with a package manager called conda used to install and update additional packages.
Research Project: Best Practices
It is a good idea to keep a set of related data, analyses, and text in a single folder. All scripts and text files within this folder can then use relative paths to the data files. Working this way makes it a lot easier to move around your project and share it with others.
Organizing your working directory
Using a consistent folder structure across your projects will help you keep things organized, and will also make it easy to find/file things in the future. This can be especially helpful when you have multiple projects. In general, you may wish to create separate directories for your scripts, data, and documents.
data/: Use this folder to store your raw data. For the sake of transparency and provenance, you should always keep a copy of your raw data. If you need to cleanup data, do it programmatically (i.e. with scripts) and make sure to separate cleaned up data from the raw data. For example, you can store raw data in files./data/raw/and clean data in./data/clean/.documents/: Use this folder to store outlines, drafts, and other text.code/: Use this folder to store your (Python) scripts for data cleaning, analysis, and plotting that you use in this particular project.
You may need to create additional directories depending on your
project needs, but these should form the backbone of your project’s
directory. For this workshop, we will need a data/ folder
to store our raw data, and we will later create a
data_output/ folder when we learn how to export data as CSV
files.
What is Programming and Coding?
Programming is the process of writing “programs” that a computer can execute and produce some (useful) output. Programming is a multi-step process that involves the following steps:
- Identifying the aspects of the real-world problem that can be solved computationally
- Identifying (the best) computational solution
- Implementing the solution in a specific computer language
- Testing, validating, and adjusting implemented solution.
While “Programming” refers to all of the above steps, “Coding” refers to step 3 only: “Implementing the solution in a specific computer language”. It’s important to note that “the best” computational solution must consider factors beyond the computer. Who is using the program, what resources/funds does your team have for this project, and the available timeline all shape and mold what “best” may be.
If you are working with Jupyter notebook:
You can type Python code into a code cell and then execute the code
by pressing Shift+Return. Output will be printed
directly under the input cell. You can recognise a code cell by the
In[ ]: at the beginning of the cell and output by
Out[ ]:. Pressing the + button in the menu
bar will add a new cell. All your commands as well as any output will be
saved with the notebook.
If you are working with Spyder:
You can either use the console or use script files (plain text files that contain your code). The console pane (in Spyder, the bottom right panel) is the place where commands written in the Python language can be typed and executed immediately by the computer. It is also where the results will be shown. You can execute commands directly in the console by pressing Return, but they will be “lost” when you close the session. Spyder uses the IPython console by default.
Since we want our code and workflow to be reproducible, it is better to type the commands in the script editor, and save them as a script. This way, there is a complete record of what we did, and anyone (including our future selves!) has an easier time reproducing the results on their computer.
Spyder allows you to execute commands directly from the script editor by using the run buttons on top. To run the entire script click Run file or press F5, to run the current line click Run selection or current line or press F9, other run buttons allow to run script cells or go into debug mode. When using F9, the command on the current line in the script (indicated by the cursor) or all of the commands in the currently selected text will be sent to the console and executed.
At some point in your analysis you may want to check the content of a variable or the structure of an object, without necessarily keeping a record of it in your script. You can type these commands and execute them directly in the console. Spyder provides the Ctrl+Shift+E and Ctrl+Shift+I shortcuts to allow you to jump between the script and the console panes.
If Python is ready to accept commands, the IPython console shows an
In [..]: prompt with the current console line number in
[]. If it receives a command (by typing, copy-pasting or
sent from the script editor), Python will execute it, display the
results in the Out [..]: cell, and come back with a new
In [..]: prompt waiting for new commands.
If Python is still waiting for you to enter more data because it
isn’t complete yet, the console will show a ...: prompt. It
means that you haven’t finished entering a complete command. This can be
because you have not typed a closing parenthesis (),
], or }) or quotation mark. When this happens,
and you thought you finished typing your command, click inside the
console window and press Esc; this will cancel the incomplete
command and return you to the In [..]: prompt.
How to learn more after the workshop?
The material we cover during this workshop will give you an initial taste of how you can use Python to analyze data for your own research. However, you will need to learn more to do advanced operations such as cleaning your dataset, using statistical methods, or creating beautiful graphics. The best way to become proficient and efficient at python, as with any other tool, is to use it to address your actual research questions. As a beginner, it can feel daunting to have to write a script from scratch, and given that many people make their code available online, modifying existing code to suit your purpose might make it easier for you to get started.
Seeking help
- check under the Help menu
- type
help() - type
?objectorhelp(object)to get information about an object - Python documentation
- Pandas documentation
Finally, a generic Google or internet search “Python task” will often either send you to the appropriate module documentation or a helpful forum where someone else has already asked your question.
I am stuck… I get an error message that I don’t understand. Start by googling the error message. However, this doesn’t always work very well, because often, package developers rely on the error catching provided by Python. You end up with general error messages that might not be very helpful to diagnose a problem (e.g. “subscript out of bounds”). If the message is very generic, you might also include the name of the function or package you’re using in your query.
However, you should check Stack Overflow. Search using the
[python] tag. Most questions have already been answered,
but the challenge is to use the right words in the search to find the
answers: https://stackoverflow.com/questions/tagged/python?tab=Votes
Asking for help
The key to receiving help from someone is for them to rapidly grasp your problem. You should make it as easy as possible to pinpoint where the issue might be.
Try to use the correct words to describe your problem. For instance, a package is not the same thing as a library. Most people will understand what you meant, but others have really strong feelings about the difference in meaning. The key point is that it can make things confusing for people trying to help you. Be as precise as possible when describing your problem.
If possible, try to reduce what doesn’t work to a simple reproducible example. If you can reproduce the problem using a very small data frame instead of your 50,000 rows and 10,000 columns one, provide the small one with the description of your problem. When appropriate, try to generalize what you are doing so even people who are not in your field can understand the question. For instance, instead of using a subset of your real dataset, create a small (3 columns, 5 rows) generic one.
Where to ask for help?
- The person sitting next to you during the workshop. Don’t hesitate to talk to your neighbor during the workshop, compare your answers, and ask for help. You might also be interested in organizing regular meetings following the workshop to keep learning from each other.
- Your friendly colleagues: if you know someone with more experience than you, they might be able and willing to help you.
- Stack Overflow: if your question hasn’t been answered before and is well crafted, chances are you will get an answer in less than 5 min. Remember to follow their guidelines on how to ask a good question.
- Python mailing lists
More resources
Key Points
- Python is an open source and platform independent programming language.
- Jupyter Notebook and the Spyder IDE are great tools to code in and interact with Python. With the large Python community it is easy to find help on the internet.
Content from Short Introduction to Programming in Python
Last updated on 2023-06-13 | Edit this page
Estimated time: 35 minutes
Overview
Questions
- How do I program in Python?
- How can I represent my data in Python?
Objectives
- Describe the advantages of using programming vs. completing repetitive tasks by hand.
- Define the following data types in Python: strings, integers, and floats.
- Perform mathematical operations in Python using basic operators.
- Define the following as it relates to Python: lists, tuples, and dictionaries.
Interpreter
Python is an interpreted language which can be used in two ways:
- “Interactively”: when you use it as an “advanced calculator”
executing one command at a time. To start Python in this mode, execute
pythonon the command line:
OUTPUT
Python 3.5.1 (default, Oct 23 2015, 18:05:06)
[GCC 4.8.3] on linux2
Type "help", "copyright", "credits" or "license" for more information.
>>>Chevrons >>> indicate an interactive prompt in
Python, meaning that it is waiting for your input.
OUTPUT
4OUTPUT
Hello World- “Scripting” Mode: executing a series of “commands” saved in text
file, usually with a
.pyextension after the name of your file:
OUTPUT
Hello WorldIntroduction to variables in Python
Assigning values to variables
One of the most basic things we can do in Python is assign values to variables:
PYTHON
text = "Data Carpentry" # An example of assigning a value to a new text variable,
# also known as a string data type in Python
number = 42 # An example of assigning a numeric value, or an integer data type
pi_value = 3.1415 # An example of assigning a floating point value (the float data type)Here we’ve assigned data to the variables text,
number and pi_value, using the assignment
operator =. To review the value of a variable, we can type
the name of the variable into the interpreter and press
Return:
OUTPUT
"Data Carpentry"Everything in Python has a type. To get the type of something, we can
pass it to the built-in function type:
OUTPUT
<class 'str'>OUTPUT
<class 'int'>OUTPUT
<class 'float'>The variable text is of type str, short for
“string”. Strings hold sequences of characters, which can be letters,
numbers, punctuation or more exotic forms of text (even emoji!).
We can also see the value of something using another built-in
function, print:
OUTPUT
Data CarpentryOUTPUT
42This may seem redundant, but in fact it’s the only way to display output in a script:
example.py
PYTHON
# A Python script file
# Comments in Python start with #
# The next line assigns the string "Data Carpentry" to the variable "text".
text = "Data Carpentry"
# The next line does nothing!
text
# The next line uses the print function to print out the value we assigned to "text"
print(text)Running the script
OUTPUT
Data CarpentryNotice that “Data Carpentry” is printed only once.
Tip: print and type are
built-in functions in Python. Later in this lesson, we will introduce
methods and user-defined functions. The Python documentation is
excellent for reference on the differences between them.
Tip: When editing scripts like example.py, be careful not to use word processors such as MS Word, as they may introduce extra information that confuses Python. In this lesson we will be using either Jupyter notebooks or the Spyder IDE, and for your everday work you may also choose any text editor such as Notepad++, VSCode, Vim, or Emacs.
Operators
We can perform mathematical calculations in Python using the basic
operators +, -, /, *, %:
OUTPUT
4OUTPUT
42OUTPUT
65536OUTPUT
3We can also use comparison and logic operators:
<, >, ==, !=, <=, >= and statements of identity
such as and, or, not. The data type returned by this is
called a boolean.
OUTPUT
FalseOUTPUT
TrueOUTPUT
TrueOUTPUT
FalseSequences: Lists and Tuples
Lists
Lists are a common data structure to hold an ordered sequence of elements. Each element can be accessed by an index. Note that Python indexes start with 0 instead of 1:
OUTPUT
1A for loop can be used to access the elements in a list
or other Python data structure one at a time:
OUTPUT
1
2
3Indentation is very important in Python. Note that
the second line in the example above is indented. Just like three
chevrons >>> indicate an interactive prompt in
Python, the three dots ... are Python’s prompt for multiple
lines. This is Python’s way of marking a block of code. [Note: you do
not type >>> or ....]
To add elements to the end of a list, we can use the
append method. Methods are a way to interact with an object
(a list, for example). We can invoke a method using the dot
. followed by the method name and a list of arguments in
parentheses. Let’s look at an example using append:
OUTPUT
[1, 2, 3, 4]To find out what methods are available for an object, we can use the
built-in help command:
OUTPUT
help(numbers)
Help on list object:
class list(object)
| list() -> new empty list
| list(iterable) -> new list initialized from iterable's items
...Tuples
A tuple is similar to a list in that it’s an ordered
sequence of elements. However, tuples can not be changed once created
(they are “immutable”). Tuples are created by placing comma-separated
values inside parentheses ().
PYTHON
# Tuples use parentheses
a_tuple = (1, 2, 3)
another_tuple = ('blue', 'green', 'red')
# Note: lists use square brackets
a_list = [1, 2, 3]Tuples vs. Lists
- What happens when you execute
a_list[1] = 5? - What happens when you execute
a_tuple[2] = 5? - What does
type(a_tuple)tell you abouta_tuple? - What information does the built-in function
len()provide? Does it provide the same information on both tuples and lists? Does thehelp()function confirm this?
- What happens when you execute
a_list[1] = 5?
The second value in a_list is replaced with
5.
- What happens when you execute
a_tuple[2] = 5?
ERROR
TypeError: 'tuple' object does not support item assignmentAs a tuple is immutable, it does not support item assignment. Elements in a list can be altered individually.
- What does
type(a_tuple)tell you abouta_tuple?
OUTPUT
<class 'tuple'>The function tells you that the variable a_tuple is an
object of the class tuple.
- What information does the built-in function
len()provide? Does it provide the same information on both tuples and lists? Does thehelp()function confirm this?
OUTPUT
3OUTPUT
3len() tells us the length of an object. It works the
same for both lists and tuples, providing us with the number of entries
in each case.
OUTPUT
Help on built-in function len in module builtins:
len(obj, /)
Return the number of items in a container.Lists and tuples are both types of container i.e. objects that can contain multiple items, the key difference being that lists are mutable i.e. they can be modified after they have been created, while tuples are not: their value cannot be modified, only overwritten.
Dictionaries
A dictionary is a container that holds pairs of objects - keys and values.
OUTPUT
'first'Dictionaries work a lot like lists - except that you index them with keys. You can think about a key as a name or unique identifier for the value it corresponds to.
OUTPUT
'one'To add an item to the dictionary we assign a value to a new key:
OUTPUT
{'first': 'one', 'second': 'two', 'third': 'three'}Using for loops with dictionaries is a little more
complicated. We can do this in two ways:
OUTPUT
'first' -> one
'second' -> two
'third' -> threeor
OUTPUT
'first' -> one
'second' -> two
'third' -> threeChanging dictionaries
- First, print the value of the
revdictionary to the screen. - Reassign the value that corresponds to the key
secondso that it no longer reads “two” but instead2. - Print the value of
revto the screen again to see if the value has changed.
Functions
Defining a section of code as a function in Python
is done using the def keyword. For example a function that
takes two arguments and returns their sum can be defined as:
OUTPUT
42Key Points
- Python is an interpreted language which can be used interactively (executing one command at a time) or in scripting mode (executing a series of commands saved in file).
- One can assign a value to a variable in Python. Those variables can be of several types, such as string, integer, floating point and complex numbers.
- Lists and tuples are similar in that they are ordered lists of elements; they differ in that a tuple is immutable (cannot be changed).
- Dictionaries are data structures that provide mappings between keys and values.
Content from Starting With Data
Last updated on 2023-05-29 | Edit this page
Estimated time: 60 minutes
Overview
Questions
- How can I import data in Python?
- What is Pandas?
- Why should I use Pandas to work with data?
Objectives
- Navigate the workshop directory and download a dataset.
- Explain what a library is and what libraries are used for.
- Describe what the Python Data Analysis Library (Pandas) is.
- Load the Python Data Analysis Library (Pandas).
- Read tabular data into Python using Pandas.
- Describe what a DataFrame is in Python.
- Access and summarize data stored in a DataFrame.
- Define indexing as it relates to data structures.
- Perform basic mathematical operations and summary statistics on data in a Pandas DataFrame.
- Create simple plots.
Working With Pandas DataFrames in Python
We can automate the process of performing data manipulations in Python. It’s efficient to spend time building the code to perform these tasks because once it’s built, we can use it over and over on different datasets that use a similar format. This makes our data manipulation processes reproducible. We can also share our code with colleagues and they can replicate the same analysis starting with the same original data.
Starting in the same spot
To help the lesson run smoothly, let’s ensure everyone is in the same directory. This should help us avoid path and file name issues. At this time please navigate to the workshop directory. If you are working in Jupyter Notebook be sure that you start your notebook in the workshop directory.
Our Data
For this lesson, we will be using the Portal Teaching data, a subset of the data from Ernst et al. Long-term monitoring and experimental manipulation of a Chihuahuan Desert ecosystem near Portal, Arizona, USA.
We will be using files from the Portal
Project Teaching Database. This section will use the
surveys.csv file that can be downloaded here: https://ndownloader.figshare.com/files/2292172
We are studying the species and
weight of animals caught in sites in our study area.
The dataset is stored as a .csv file: each row holds
information for a single animal, and the columns represent:
| Column | Description |
|---|---|
| record_id | Unique id for the observation |
| month | month of observation |
| day | day of observation |
| year | year of observation |
| plot_id | ID of a particular site |
| species_id | 2-letter code |
| sex | sex of animal (“M”, “F”) |
| hindfoot_length | length of the hindfoot in mm |
| weight | weight of the animal in grams |
The first few rows of our first file look like this:
OUTPUT
record_id,month,day,year,plot_id,species_id,sex,hindfoot_length,weight
1,7,16,1977,2,NL,M,32,
2,7,16,1977,3,NL,M,33,
3,7,16,1977,2,DM,F,37,
4,7,16,1977,7,DM,M,36,
5,7,16,1977,3,DM,M,35,
6,7,16,1977,1,PF,M,14,
7,7,16,1977,2,PE,F,,
8,7,16,1977,1,DM,M,37,
9,7,16,1977,1,DM,F,34,About Libraries
A library in Python contains a set of tools (called functions) that perform tasks on our data. Importing a library is like getting a piece of lab equipment out of a storage locker and setting it up on the bench for use in a project. Once a library is set up, it can be used or called to perform the task(s) it was built to do.
Pandas in Python
One of the best options for working with tabular data in Python is to use the Python Data Analysis Library (a.k.a. Pandas). The Pandas library provides data structures, produces high quality plots with matplotlib and integrates nicely with other libraries that use NumPy (which is another Python library) arrays.
Python doesn’t load all of the libraries available to it by default.
We have to add an import statement to our code in order to
use library functions. To import a library, we use the syntax
import libraryName. If we want to give the library a
nickname to shorten the command, we can add
as nickNameHere. An example of importing the pandas library
using the common nickname pd is below.
Each time we call a function that’s in a library, we use the syntax
LibraryName.FunctionName. Adding the library name with a
. before the function name tells Python where to find the
function. In the example above, we have imported Pandas as
pd. This means we don’t have to type out
pandas each time we call a Pandas function.
Reading CSV Data Using Pandas
We will begin by locating and reading our survey data which are in
CSV format. CSV stands for Comma-Separated Values and is a common way to
store formatted data. Other symbols may also be used, so you might see
tab-separated, colon-separated or space separated files. pandas can work
with each of these types of separators, as it allows you to specify the
appropriate separator for your data. CSV files (and other -separated
value file types) make it easy to share data, and can be imported and
exported from many applications, including Microsoft Excel. For more
details on CSV files, see the Data
Organisation in Spreadsheets lesson. We can use Pandas’
read_csv function to pull the file directly into a DataFrame.
So What’s a DataFrame?
A DataFrame is a 2-dimensional data structure that can store data of
different types (including strings, numbers, categories and more) in
columns. It is similar to a spreadsheet or an SQL table or the
data.frame in R. A DataFrame always has an index (0-based).
An index refers to the position of an element in the data structure.
PYTHON
# Note that pd.read_csv is used because we imported pandas as pd
pd.read_csv("data/surveys.csv")The above command yields the output below:
OUTPUT
record_id month day year plot_id species_id sex hindfoot_length weight
0 1 7 16 1977 2 NL M 32.0 NaN
1 2 7 16 1977 3 NL M 33.0 NaN
2 3 7 16 1977 2 DM F 37.0 NaN
3 4 7 16 1977 7 DM M 36.0 NaN
4 5 7 16 1977 3 DM M 35.0 NaN
... ... ... ... ... ... ... ... ... ...
35544 35545 12 31 2002 15 AH NaN NaN NaN
35545 35546 12 31 2002 15 AH NaN NaN NaN
35546 35547 12 31 2002 10 RM F 15.0 14.0
35547 35548 12 31 2002 7 DO M 36.0 51.0
35548 35549 12 31 2002 5 NaN NaN NaN NaN
[35549 rows x 9 columns]We can see that there were 35,549 rows parsed. Each row has 9
columns. The first column is the index of the DataFrame. The index is
used to identify the position of the data, but it is not an actual
column of the DataFrame. It looks like the read_csv
function in Pandas read our file properly. However, we haven’t saved any
data to memory so we can work with it. We need to assign the DataFrame
to a variable. Remember that a variable is a name for a value, such as
x, or data. We can create a new object with a
variable name by assigning a value to it using =.
Let’s call the imported survey data surveys_df:
Note that Python does not produce any output on the screen when you
assign the imported DataFrame to a variable. We can view the value of
the surveys_df object by typing its name into the Python
command prompt.
which prints contents like above.
Note: if the output is too wide to print on your narrow terminal window, you may see something slightly different as the large set of data scrolls past. You may see simply the last column of data:
OUTPUT
17 NaN
18 NaN
19 NaN
20 NaN
21 NaN
22 NaN
23 NaN
24 NaN
25 NaN
26 NaN
27 NaN
28 NaN
29 NaN
... ...
35519 36.0
35520 48.0
35521 45.0
35522 44.0
35523 27.0
35524 26.0
35525 24.0
35526 43.0
35527 NaN
35528 25.0
35529 NaN
35530 NaN
35531 43.0
35532 48.0
35533 56.0
35534 53.0
35535 42.0
35536 46.0
35537 31.0
35538 68.0
35539 23.0
35540 31.0
35541 29.0
35542 34.0
35543 NaN
35544 NaN
35545 NaN
35546 14.0
35547 51.0
35548 NaN
[35549 rows x 9 columns]Don’t worry: all the data is there! You can confirm this by scrolling
upwards, or by looking at the [# of rows x # of columns]
block at the end of the output.
You can also use surveys_df.head() to view only the
first few rows of the dataset in an output that is easier to fit in one
window. After doing this, you can see that pandas has neatly formatted
the data to fit our screen:
PYTHON
surveys_df.head() # The head() method displays the first several lines of a file. It
# is discussed below.OUTPUT
record_id month day year plot_id species_id sex hindfoot_length \
5 6 7 16 1977 1 PF M 14.0
6 7 7 16 1977 2 PE F NaN
7 8 7 16 1977 1 DM M 37.0
8 9 7 16 1977 1 DM F 34.0
9 10 7 16 1977 6 PF F 20.0
weight
5 NaN
6 NaN
7 NaN
8 NaN
9 NaNExploring Our Species Survey Data
Again, we can use the type function to see what kind of
thing surveys_df is:
OUTPUT
<class 'pandas.core.frame.DataFrame'>As expected, it’s a DataFrame (or, to use the full name that Python
uses to refer to it internally, a
pandas.core.frame.DataFrame).
What kind of things does surveys_df contain? DataFrames
have an attribute called dtypes that answers this:
OUTPUT
record_id int64
month int64
day int64
year int64
plot_id int64
species_id object
sex object
hindfoot_length float64
weight float64
dtype: objectAll the values in a single column have the same type. For example,
values in the month column have type int64, which is a kind
of integer. Cells in the month column cannot have fractional values, but
values in weight and hindfoot_length columns can, because they have type
float64. The object type doesn’t have a very
helpful name, but in this case it represents strings (such as ‘M’ and
‘F’ in the case of sex).
We’ll talk a bit more about what the different formats mean in a different lesson.
Useful Ways to View DataFrame Objects in Python
There are many ways to summarize and access the data stored in DataFrames, using attributes and methods provided by the DataFrame object.
Attributes are features of an object. For example, the
shape attribute will output the size (the number of rows
and columns) of an object. To access an attribute, use the DataFrame
object name followed by the attribute name
df_object.attribute. For example, using the DataFrame
surveys_df and attribute columns, an index of
all the column names in the DataFrame can be accessed with
surveys_df.columns.
Methods are like functions, but they only work on particular kinds of
objects. As an example, the head() method
works on DataFrames. Methods are called in a similar fashion to
attributes, using the syntax df_object.method(). Using
surveys_df.head() gets the first few rows in the DataFrame
surveys_df using the head() method. With a
method, we can supply extra information in the parentheses to control
behaviour.
Let’s look at the data using these.
Challenge - DataFrames
Using our DataFrame surveys_df, try out the
attributes & methods below to see
what they return.
surveys_df.columnssurveys_df.shapeTake note of the output ofshape- what format does it return the shape of the DataFrame in?
HINT: More on tuples, here.
surveys_df.head()Also, what doessurveys_df.head(15)do?surveys_df.tail()
-
surveys_df.columnsprovides the names of the columns in the DataFrame. -
surveys_df.shapeprovides the dimensions of the DataFrame as a tuple in(r,c)format, whereris the number of rows andcthe number of columns. -
surveys_df.head()returns the first 5 lines of the DataFrame, annotated with column and row labels. Adding an integer as an argument to the function specifies the number of lines to display from the top of the DataFrame, e.g.surveys_df.head(15)will return the first 15 lines. -
surveys_df.tail()will display the last 5 lines, and behaves similarly to thehead()method.
Working through solutions to the challenge above can provide a good
opportunity to recap about mutability and immutability of different
objects. Show that the DataFrame index ( the columns
attribute) is immutable, e.g.
surveys_df.columns[4] = "plotid" returns a
TypeError.
Adapting the name is done with the rename function:
Calculating Statistics From Data In A Pandas DataFrame
We’ve read our data into Python. Next, let’s perform some quick summary statistics to learn more about the data that we’re working with. We might want to know how many animals were collected in each site, or how many of each species were caught. We can perform summary stats quickly using groups. But first we need to figure out what we want to group by.
Let’s begin by exploring our data:
which returns:
OUTPUT
Index(['record_id', 'month', 'day', 'year', 'plot_id', 'species_id', 'sex',
'hindfoot_length', 'weight'],
dtype='object')Let’s get a list of all the species. The pd.unique
function tells us all of the unique values in the
species_id column.
which returns:
OUTPUT
array(['NL', 'DM', 'PF', 'PE', 'DS', 'PP', 'SH', 'OT', 'DO', 'OX', 'SS',
'OL', 'RM', nan, 'SA', 'PM', 'AH', 'DX', 'AB', 'CB', 'CM', 'CQ',
'RF', 'PC', 'PG', 'PH', 'PU', 'CV', 'UR', 'UP', 'ZL', 'UL', 'CS',
'SC', 'BA', 'SF', 'RO', 'AS', 'SO', 'PI', 'ST', 'CU', 'SU', 'RX',
'PB', 'PL', 'PX', 'CT', 'US'], dtype=object)Challenge - Statistics
Create a list of unique site IDs (“plot_id”) found in the surveys data. Call it
site_names. How many unique sites are there in the data? How many unique species are in the data?What is the difference between
len(site_names)andsurveys_df['plot_id'].nunique()?
site_names = pd.unique(surveys_df["plot_id"])
- How many unique sites are in the data?
site_names.sizeorlen(site_names)provide the answer: 24 - How many unique species are in the data?
len(pd.unique(surveys_df["species_id"]))tells us there are 49 species
-
len(site_names)andsurveys_df['plot_id'].nunique()both provide the same output: they are alternative ways of getting the unique values. Thenuniquemethod combines the count and unique value extraction, and can help avoid the creation of intermediate variables likesite_names.
Groups in Pandas
We often want to calculate summary statistics grouped by subsets or attributes within fields of our data. For example, we might want to calculate the average weight of all individuals per site.
We can calculate basic statistics for all records in a single column using the syntax below:
gives output
PYTHON
count 32283.000000
mean 42.672428
std 36.631259
min 4.000000
25% 20.000000
50% 37.000000
75% 48.000000
max 280.000000
Name: weight, dtype: float64In pandas prior to version 0.18.1 there is a bug causing
surveys_df['weight'].describe() to return a runtime
error.
We can also extract one specific metric if we wish:
PYTHON
surveys_df['weight'].min()
surveys_df['weight'].max()
surveys_df['weight'].mean()
surveys_df['weight'].std()
surveys_df['weight'].count()But if we want to summarize by one or more variables, for example
sex, we can use Pandas’ .groupby method.
Once we’ve created a groupby DataFrame, we can quickly calculate summary
statistics by a group of our choice.
The pandas function describe will
return descriptive stats including: mean, median, max, min, std and
count for a particular column in the data. Pandas’ describe
function will only return summary values for columns containing numeric
data.
PYTHON
# Summary statistics for all numeric columns by sex
grouped_data.describe()
# Provide the mean for each numeric column by sex
grouped_data.mean(numeric_only=True)grouped_data.mean(numeric_only=True)
OUTPUT:
OUTPUT
record_id month day year plot_id \
sex
F 18036.412046 6.583047 16.007138 1990.644997 11.440854
M 17754.835601 6.392668 16.184286 1990.480401 11.098282
hindfoot_length weight
sex
F 28.836780 42.170555
M 29.709578 42.995379
The groupby command is powerful in that it allows us to
quickly generate summary stats.
Challenge - Summary Data
- How many recorded individuals are female
Fand how many maleM? - What happens when you group by two columns using the following syntax and then calculate mean values?
grouped_data2 = surveys_df.groupby(['plot_id', 'sex'])grouped_data2.mean(numeric_only=True)
- Summarize weight values for each site in your data. HINT: you can
use the following syntax to only create summary statistics for one
column in your data.
by_site['weight'].describe()
- The first column of output from
grouped_data.describe()(count) tells us that the data contains 15690 records for female individuals and 17348 records for male individuals.- Note that these two numbers do not sum to 35549, the total number of
rows we know to be in the
surveys_dfDataFrame. Why do you think some records were excluded from the grouping?
- Note that these two numbers do not sum to 35549, the total number of
rows we know to be in the
- Calling the
mean()method on data grouped by these two columns calculates and returns the mean value for each combination of plot and sex.- Note that the mean is not meaningful for some variables, e.g. day,
month, and year. You can specify particular columns and particular
summary statistics using the
agg()method (short for aggregate), e.g. to obtain the last survey year, median foot-length and mean weight for each plot/sex combination:
- Note that the mean is not meaningful for some variables, e.g. day,
month, and year. You can specify particular columns and particular
summary statistics using the
PYTHON
surveys_df.groupby(['plot_id', 'sex']).agg({"year": 'max',
"hindfoot_length": 'median',
"weight": 'mean'})surveys_df.groupby(['plot_id'])['weight'].describe()
OUTPUT
count mean std min 25% 50% 75% max
plot_id
1 1903.0 51.822911 38.176670 4.0 30.0 44.0 53.0 231.0
2 2074.0 52.251688 46.503602 5.0 24.0 41.0 50.0 278.0
3 1710.0 32.654386 35.641630 4.0 14.0 23.0 36.0 250.0
4 1866.0 47.928189 32.886598 4.0 30.0 43.0 50.0 200.0
5 1092.0 40.947802 34.086616 5.0 21.0 37.0 48.0 248.0
6 1463.0 36.738893 30.648310 5.0 18.0 30.0 45.0 243.0
7 638.0 20.663009 21.315325 4.0 11.0 17.0 23.0 235.0
8 1781.0 47.758001 33.192194 5.0 26.0 44.0 51.0 178.0
9 1811.0 51.432358 33.724726 6.0 36.0 45.0 50.0 275.0
10 279.0 18.541219 20.290806 4.0 10.0 12.0 21.0 237.0
11 1793.0 43.451757 28.975514 5.0 26.0 42.0 48.0 212.0
12 2219.0 49.496169 41.630035 6.0 26.0 42.0 50.0 280.0
13 1371.0 40.445660 34.042767 5.0 20.5 33.0 45.0 241.0
14 1728.0 46.277199 27.570389 5.0 36.0 44.0 49.0 222.0
15 869.0 27.042578 35.178142 4.0 11.0 18.0 26.0 259.0
16 480.0 24.585417 17.682334 4.0 12.0 20.0 34.0 158.0
17 1893.0 47.889593 35.802399 4.0 27.0 42.0 50.0 216.0
18 1351.0 40.005922 38.480856 5.0 17.5 30.0 44.0 256.0
19 1084.0 21.105166 13.269840 4.0 11.0 19.0 27.0 139.0
20 1222.0 48.665303 50.111539 5.0 17.0 31.0 47.0 223.0
21 1029.0 24.627794 21.199819 4.0 10.0 22.0 31.0 190.0
22 1298.0 54.146379 38.743967 5.0 29.0 42.0 54.0 212.0
23 369.0 19.634146 18.382678 4.0 10.0 14.0 23.0 199.0
24 960.0 43.679167 45.936588 4.0 19.0 27.5 45.0 251.0Quickly Creating Summary Counts in Pandas
Let’s next count the number of samples for each species. We can do
this in a few ways, but we’ll use groupby combined with
a count() method.
PYTHON
# Count the number of samples by species
species_counts = surveys_df.groupby('species_id')['record_id'].count()
print(species_counts)Or, we can also count just the rows that have the species “DO”:
Challenge - Make a list
What’s another way to create a list of species and associated
count of the records in the data? Hint: you can perform
count, min, etc. functions on groupby
DataFrames in the same way you can perform them on regular
DataFrames.
As well as calling count() on the record_id
column of the grouped DataFrame as above, an equivalent result can be
obtained by extracting record_id from the result of
count() called directly on the grouped DataFrame:
OUTPUT
species_id
AB 303
AH 437
AS 2
BA 46
CB 50
CM 13
CQ 16
CS 1
CT 1
CU 1
CV 1
DM 10596
DO 3027
DS 2504
DX 40
NL 1252
OL 1006
OT 2249
OX 12
PB 2891
PC 39
PE 1299
PF 1597
PG 8
PH 32
PI 9
PL 36
PM 899
PP 3123
PU 5
PX 6
RF 75
RM 2609
RO 8
RX 2
SA 75
SC 1
SF 43
SH 147
SO 43
SS 248
ST 1
SU 5
UL 4
UP 8
UR 10
US 4
ZL 2Basic Math Functions
If we wanted to, we could apply a mathmatical operation like addition or division on an entire column of our data. For example, let’s multiply all weight values by 2.
A more practical use of this might be to normalize the data according to a mean, area, or some other value calculated from our data.
Quick & Easy Plotting Data Using Pandas
We can plot our summary stats using Pandas, too.
PYTHON
# Make sure figures appear inline in Ipython Notebook
%matplotlib inline
# Create a quick bar chart
species_counts.plot(kind='bar');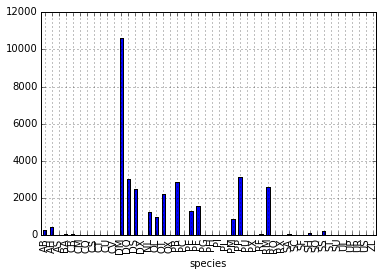 Count per species site
Count per species site
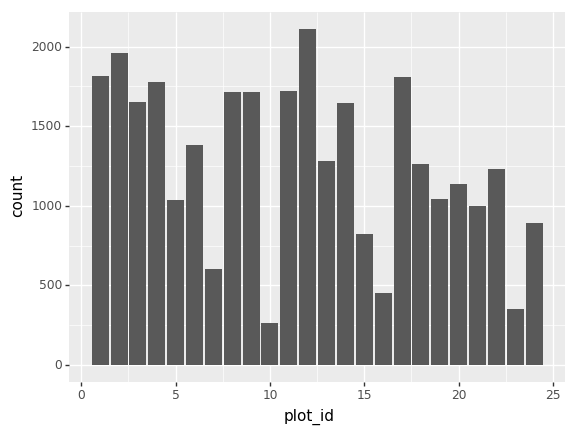
We can also look at how many animals were captured in each site:
PYTHON
total_count = surveys_df.groupby('plot_id')['record_id'].nunique()
# Let's plot that too
total_count.plot(kind='bar');Challenge - Plots
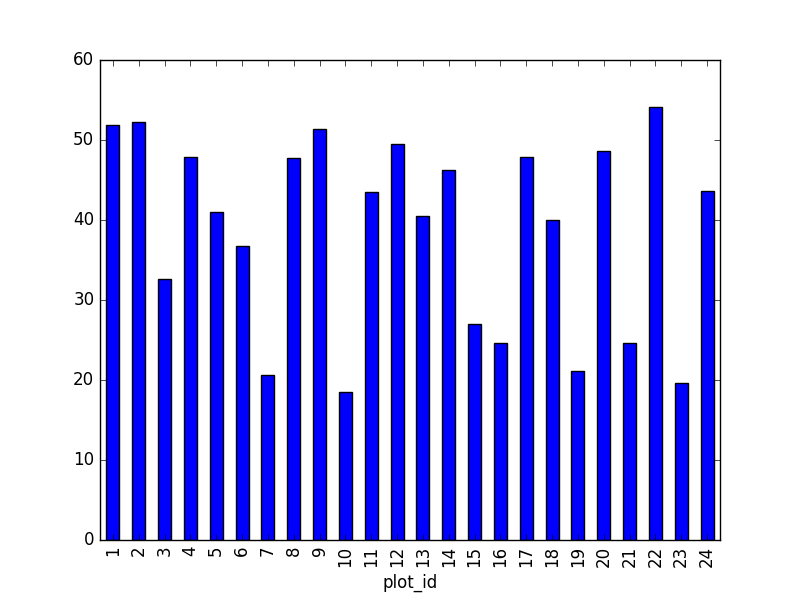
- Create a plot of average weight across all species per site.
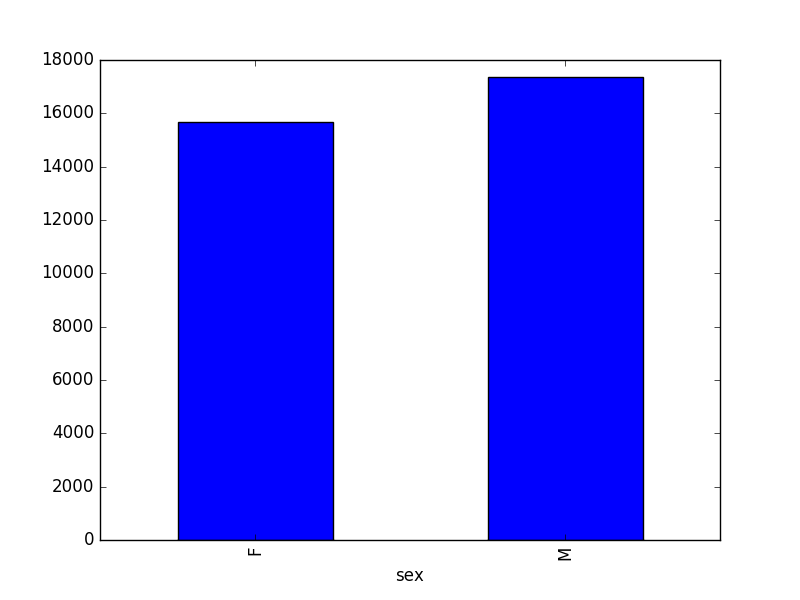
- Create a plot of total males versus total females for the entire dataset.
surveys_df.groupby('plot_id').mean()["weight"].plot(kind='bar')
surveys_df.groupby('sex').count()["record_id"].plot(kind='bar')
Summary Plotting Challenge
Create a stacked bar plot, with weight on the Y axis, and the stacked variable being sex. The plot should show total weight by sex for each site. Some tips are below to help you solve this challenge:
- For more information on pandas plots, see pandas’ documentation page on visualization.
- You can use the code that follows to create a stacked bar plot but the data to stack need to be in individual columns. Here’s a simple example with some data where ‘a’, ‘b’, and ‘c’ are the groups, and ‘one’ and ‘two’ are the subgroups.
PYTHON
d = {'one' : pd.Series([1., 2., 3.], index=['a', 'b', 'c']), 'two' : pd.Series([1., 2., 3., 4.], index=['a', 'b', 'c', 'd'])}
pd.DataFrame(d)shows the following data
OUTPUT
one two
a 1 1
b 2 2
c 3 3
d NaN 4We can plot the above with
PYTHON
# Plot stacked data so columns 'one' and 'two' are stacked
my_df = pd.DataFrame(d)
my_df.plot(kind='bar', stacked=True, title="The title of my graph")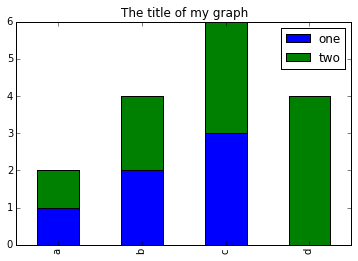
- You can use the
.unstack()method to transform grouped data into columns for each plotting. Try running.unstack()on some DataFrames above and see what it yields.
Start by transforming the grouped data (by site and sex) into an unstacked layout, then create a stacked plot.
First we group data by site and by sex, and then calculate a total for each site.
PYTHON
by_site_sex = surveys_df.groupby(['plot_id', 'sex'])
site_sex_count = by_site_sex['weight'].sum()This calculates the sums of weights for each sex within each site as a table
OUTPUT
site sex
plot_id sex
1 F 38253
M 59979
2 F 50144
M 57250
3 F 27251
M 28253
4 F 39796
M 49377
<other sites removed for brevity>Below we’ll use .unstack() on our grouped data to figure
out the total weight that each sex contributed to each site.
PYTHON
by_site_sex = surveys_df.groupby(['plot_id', 'sex'])
site_sex_count = by_site_sex['weight'].sum()
site_sex_count.unstack()The unstack method above will display the following
output:
OUTPUT
sex F M
plot_id
1 38253 59979
2 50144 57250
3 27251 28253
4 39796 49377
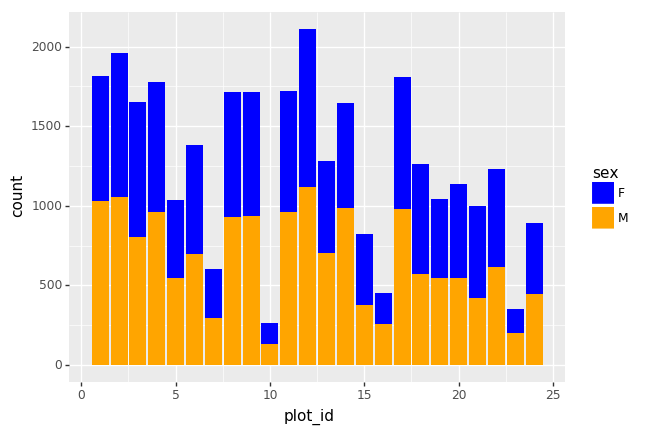
<other sites removed for brevity>Now, create a stacked bar plot with that data where the weights for each sex are stacked by site.
Rather than display it as a table, we can plot the above data by stacking the values of each sex as follows:
PYTHON
by_site_sex = surveys_df.groupby(['plot_id', 'sex'])
site_sex_count = by_site_sex['weight'].sum()
spc = site_sex_count.unstack()
s_plot = spc.plot(kind='bar', stacked=True, title="Total weight by site and sex")
s_plot.set_ylabel("Weight")
s_plot.set_xlabel("Plot")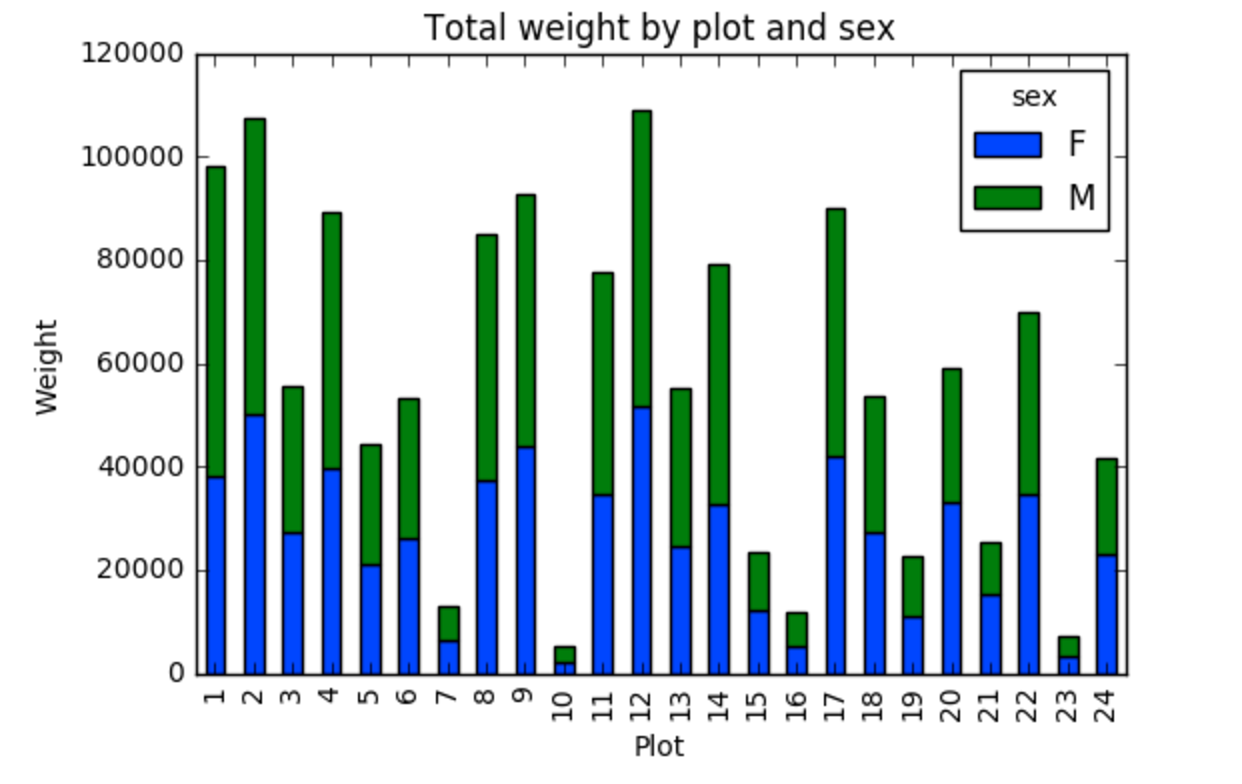
Key Points
- Libraries enable us to extend the functionality of Python.
- Pandas is a popular library for working with data.
- A Dataframe is a Pandas data structure that allows one to access data by column (name or index) or row.
- Aggregating data using the
groupby()function enables you to generate useful summaries of data quickly. - Plots can be created from DataFrames or subsets of data that have
been generated with
groupby().
Content from Indexing, Slicing and Subsetting DataFrames in Python
Last updated on 2023-08-18 | Edit this page
Estimated time: 60 minutes
Overview
Questions
- How can I access specific data within my data set?
- How can Python and Pandas help me to analyse my data?
Objectives
- Describe what 0-based indexing is.
- Manipulate and extract data using column headings and index locations.
- Employ slicing to select sets of data from a DataFrame.
- Employ label and integer-based indexing to select ranges of data in a dataframe.
- Reassign values within subsets of a DataFrame.
- Create a copy of a DataFrame.
- Query / select a subset of data using a set of criteria using the
following operators:
==,!=,>,<,>=,<=. - Locate subsets of data using masks.
- Describe BOOLEAN objects in Python and manipulate data using BOOLEANs.
Tip: use .head() method throughout this lesson to keep
your display neater for students. Encourage students to try with and
without .head() to reinforce this useful tool and then to
use it or not at their preference.
For example, if a student worries about keeping up in pace with
typing, let them know they can skip the .head(), but that
you’ll use it to keep more lines of previous steps visible.
In the first episode of this lesson, we read a CSV file into a pandas’ DataFrame. We learned how to:
- save a DataFrame to a named object,
- perform basic math on data,
- calculate summary statistics, and
- create plots based on the data we loaded into pandas.
In this lesson, we will explore ways to access different parts of the data using:
- indexing,
- slicing, and
- subsetting.
Loading our data
We will continue to use the surveys dataset that we worked with in the last episode. Let’s reopen and read in the data again:
Indexing and Slicing in Python
We often want to work with subsets of a DataFrame object. There are different ways to accomplish this including: using labels (column headings), numeric ranges, or specific x,y index locations.
Selecting data using Labels (Column Headings)
We use square brackets [] to select a subset of a Python
object. For example, we can select all data from a column named
species_id from the surveys_df DataFrame by
name. There are two ways to do this:
PYTHON
# TIP: use the .head() method we saw earlier to make output shorter
# Method 1: select a 'subset' of the data using the column name
surveys_df['species_id']
# Method 2: use the column name as an 'attribute'; gives the same output
surveys_df.species_idWe can also create a new object that contains only the data within
the species_id column as follows:
PYTHON
# Creates an object, surveys_species, that only contains the `species_id` column
surveys_species = surveys_df['species_id']We can pass a list of column names too, as an index to select columns in that order. This is useful when we need to reorganize our data.
NOTE: If a column name is not contained in the DataFrame, an exception (error) will be raised.
PYTHON
# Select the species and plot columns from the DataFrame
surveys_df[['species_id', 'plot_id']]
# What happens when you flip the order?
surveys_df[['plot_id', 'species_id']]
# What happens if you ask for a column that doesn't exist?
surveys_df['speciess']Python tells us what type of error it is in the traceback, at the
bottom it says KeyError: 'speciess' which means that
speciess is not a valid column name (nor a valid key in the
related Python data type dictionary).
Reminder
The Python language and its modules (such as Pandas) define reserved
words that should not be used as identifiers when assigning objects and
variable names. Examples of reserved words in Python include Boolean
values True and False, operators
and, or, and not, among others.
The full list of reserved words for Python version 3 is provided at https://docs.python.org/3/reference/lexical_analysis.html#identifiers.
When naming objects and variables, it’s also important to avoid using
the names of built-in data structures and methods. For example, a
list is a built-in data type. It is possible to use the word
‘list’ as an identifier for a new object, for example
list = ['apples', 'oranges', 'bananas']. However, you would
then be unable to create an empty list using list() or
convert a tuple to a list using list(sometuple).
Extracting Range based Subsets: Slicing
Reminder
Python uses 0-based indexing.
Let’s remind ourselves that Python uses 0-based indexing. This means
that the first element in an object is located at position
0. This is different from other tools like R and Matlab
that index elements within objects starting at 1.
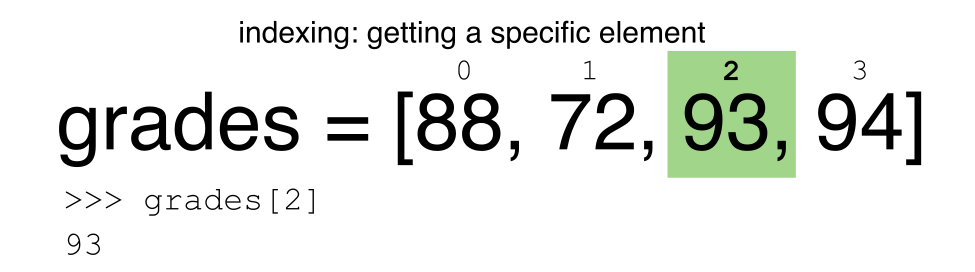
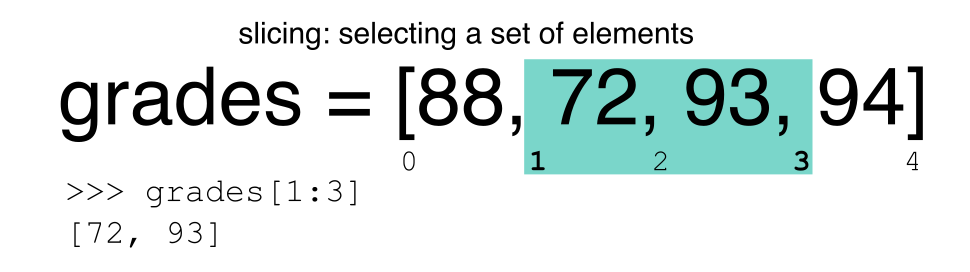
a[0]returns1, as Python starts with element 0 (this may be different from what you have previously experience with other languages e.g. MATLAB and R)a[5]raises anIndexErrorThe error is raised because the list
ahas no element with index 5: it has only five entries, indexed from 0 to 4.-
a[len(a)]also raises anIndexError.len(a)returns5, makinga[len(a)]equivalent toa[5]. To retreive the final element of a list, us the index-1, e.g.OUTPUT
5
Slicing Subsets of Rows in Python
Slicing using the [] operator selects a set of rows
and/or columns from a DataFrame. To slice out a set of rows, you use the
following syntax: data[start:stop]. When slicing in pandas
the start bound is included in the output. The stop bound is one step
BEYOND the row you want to select. So if you want to select rows 0, 1
and 2 your code would look like this:
The stop bound in Python is different from what you might be used to in languages like Matlab and R.
PYTHON
# Select the first 5 rows (rows 0, 1, 2, 3, 4)
surveys_df[:5]
# Select the last element in the list
# (the slice starts at the last element, and ends at the end of the list)
surveys_df[-1:]We can also reassign values within subsets of our DataFrame.
But before we do that, let’s look at the difference between the concept of copying objects and the concept of referencing objects in Python.
Copying Objects vs Referencing Objects in Python
Let’s start with an example:
PYTHON
# Using the 'copy() method'
true_copy_surveys_df = surveys_df.copy()
# Using the '=' operator
ref_surveys_df = surveys_dfYou might think that the code
ref_surveys_df = surveys_df creates a fresh distinct copy
of the surveys_df DataFrame object. However, using the
= operator in the simple statement y = x does
not create a copy of our DataFrame. Instead,
y = x creates a new variable y that references
the same object that x refers to. To state
this another way, there is only one object (the
DataFrame), and both x and y refer to it.
In contrast, the copy() method for a DataFrame creates a
true copy of the DataFrame.
Let’s look at what happens when we reassign the values within a subset of the DataFrame that references another DataFrame object:
PYTHON
# Assign the value `0` to the first three rows of data in the DataFrame
ref_surveys_df[0:3] = 0Let’s try the following code:
PYTHON
# ref_surveys_df was created using the '=' operator
ref_surveys_df.head()
# true_copy_surveys_df was created using the copy() function
true_copy_surveys_df.head()
# surveys_df is the original dataframe
surveys_df.head()What is the difference between these three dataframes?
When we assigned the first 3 rows the value of 0 using
the ref_surveys_df DataFrame, the surveys_df
DataFrame is modified too. Remember we created the reference
ref_surveys_df object above when we did
ref_surveys_df = surveys_df. Remember
surveys_df and ref_surveys_df refer to the
same exact DataFrame object. If either one changes the object, the other
will see the same changes to the reference object.
However - true_copy_surveys_df was created via the
copy() function. It retains the original values for the
first three rows.
To review and recap:
-
Copy uses the dataframe’s
copy()method -
A Reference is created using the
=operator
Okay, that’s enough of that. Let’s create a brand new clean dataframe from the original data CSV file.
Slicing Subsets of Rows and Columns in Python
We can select specific ranges of our data in both the row and column directions using either label or integer-based indexing.
ilocis primarily an integer based indexing counting from 0. That is, your specify rows and columns giving a number. Thus, the first row is row 0, the second column is column 1, etc.locis primarily a label based indexing where you can refer to rows and columns by their name. E.g., column ‘month’. Note that integers may be used, but they are interpreted as a label.
To select a subset of rows and columns from our
DataFrame, we can use the iloc method. For example, we can
select month, day and year (columns 2, 3 and 4 if we start counting at
1), like this:
which gives the output
OUTPUT
month day year
0 7 16 1977
1 7 16 1977
2 7 16 1977Notice that we asked for a slice from 0:3. This yielded 3 rows of data. When you ask for 0:3, you are actually telling Python to start at index 0 and select rows 0, 1, 2 up to but not including 3.
Let’s explore some other ways to index and select subsets of data:
PYTHON
# Select all columns for rows of index values 0 and 10
surveys_df.loc[[0, 10], :]
# What does this do?
surveys_df.loc[0, ['species_id', 'plot_id', 'weight']]
# What happens when you type the code below?
surveys_df.loc[[0, 10, 35549], :]NOTE: Labels must be found in the DataFrame or you
will get a KeyError.
Indexing by labels loc differs from indexing by integers
iloc. With loc, both the start bound and the
stop bound are inclusive. When using loc,
integers can be used, but the integers refer to the index label
and not the position. For example, using loc and select 1:4
will get a different result than using iloc to select rows
1:4.
We can also select a specific data value using a row and column
location within the DataFrame and iloc indexing:
In this iloc example,
gives the output
OUTPUT
'F'Remember that Python indexing begins at 0. So, the index location [2, 6] selects the element that is 3 rows down and 7 columns over in the DataFrame.
It is worth noting that rows are selected when using loc
with a single list of labels (or iloc with a single list of
integers). However, unlike loc or iloc,
indexing a data frame directly with labels will select columns (e.g.
surveys_df[['species_id', 'plot_id', 'weight']]), while
ranges of integers will select rows (e.g. surveys_df[0:13]). Direct
indexing of rows is redundant with using iloc, and will
raise a KeyError if a single integer or list is used; the
error will also occur if index labels are used without loc
(or column labels used with it). A useful rule of thumb is the
following: integer-based slicing is best done with iloc and
will avoid errors (and is generally consistent with indexing of Numpy
arrays), label-based slicing of rows is done with loc, and
slicing of columns by directly indexing column names.
Challenge - Range
- What happens when you execute:
surveys_df[0:1]surveys_df[0]surveys_df[:4]surveys_df[:-1]
- What happens when you call:
surveys_df.iloc[0:1]surveys_df.iloc[0]surveys_df.iloc[:4, :]surveys_df.iloc[0:4, 1:4]surveys_df.loc[0:4, 1:4]
- How are the last two commands different?
-
-
surveys_df[0:3]returns the first three rows of the DataFrame:
OUTPUT
record_id month day year plot_id species_id sex hindfoot_length weight 0 1 7 16 1977 2 NL M 32.0 NaN 1 2 7 16 1977 3 NL M 33.0 NaN 2 3 7 16 1977 2 DM F 37.0 NaN-
surveys_df[0]results in a ‘KeyError’, since direct indexing of a row is redundant withiloc. -
surveys_df[0:1]can be used to obtain only the first row. -
surveys_df[:5]slices from the first row to the fifth:
OUTPUT
record_id month day year plot_id species_id sex hindfoot_length weight 0 1 7 16 1977 2 NL M 32.0 NaN 1 2 7 16 1977 3 NL M 33.0 NaN 2 3 7 16 1977 2 DM F 37.0 NaN 3 4 7 16 1977 7 DM M 36.0 NaN 4 5 7 16 1977 3 DM M 35.0 NaN-
surveys_df[:-1]provides everything except the final row of the DataFrame. You can use negative index numbers to count backwards from the last entry.
-
-
surveys_df.iloc[0:1]returns the first row -
surveys_df.iloc[0]returns the first row as a named list -
surveys_df.iloc[:4, :]returns all columns of the first four rows -
surveys_df.iloc[0:4, 1:4]selects specified columns of the first four rows -
surveys_df.loc[0:4, 1:4]results in a ‘TypeError’ - see below.
-
- While
ilocuses integers as indices and slices accordingly,locworks with labels. It is like accessing values from a dictionary, asking for the key names. Column names 1:4 do not exist, so the call tolocabove results in an error. Check also the difference betweensurveys_df.loc[0:4]andsurveys_df.iloc[0:4].
Subsetting Data using Criteria
We can also select a subset of our data using criteria. For example, we can select all rows that have a year value of 2002:
Which produces the following output:
PYTHON
record_id month day year plot_id species_id sex hindfoot_length weight
33320 33321 1 12 2002 1 DM M 38 44
33321 33322 1 12 2002 1 DO M 37 58
33322 33323 1 12 2002 1 PB M 28 45
33323 33324 1 12 2002 1 AB NaN NaN NaN
33324 33325 1 12 2002 1 DO M 35 29
...
35544 35545 12 31 2002 15 AH NaN NaN NaN
35545 35546 12 31 2002 15 AH NaN NaN NaN
35546 35547 12 31 2002 10 RM F 15 14
35547 35548 12 31 2002 7 DO M 36 51
35548 35549 12 31 2002 5 NaN NaN NaN NaN
[2229 rows x 9 columns]Or we can select all rows that do not contain the year 2002:
We can define sets of criteria too:
Python Syntax Cheat Sheet
We can use the syntax below when querying data by criteria from a DataFrame. Experiment with selecting various subsets of the “surveys” data.
- Equals:
== - Not equals:
!= - Greater than, less than:
>or< - Greater than or equal to
>= - Less than or equal to
<=
Challenge - Queries
Select a subset of rows in the
surveys_dfDataFrame that contain data from the year 1999 and that contain weight values less than or equal to 8. How many rows did you end up with? What did your neighbor get?-
You can use the
isincommand in Python to query a DataFrame based upon a list of values as follows:Use the
isinfunction to find all plots that contain particular species in the “surveys” DataFrame. How many records contain these values? Experiment with other queries. Create a query that finds all rows with a weight value greater than or equal to 0.
The
~symbol in Python can be used to return the OPPOSITE of the selection that you specify. It is equivalent to is not in. Write a query that selects all rows with sex NOT equal to ‘M’ or ‘F’ in the “surveys” data.
-
OUTPUT
record_id month day year plot_id species_id sex hindfoot_length weight 29082 29083 1 16 1999 21 RM M 16.0 8.0 29196 29197 2 20 1999 18 RM M 18.0 8.0 29421 29422 3 15 1999 16 RM M 15.0 8.0 29903 29904 10 10 1999 4 PP M 20.0 7.0 29905 29906 10 10 1999 4 PP M 21.0 4.0If you are only interested in how many rows meet the criteria, the sum of
Truevalues could be used instead:OUTPUT
5 -
For example, using
PBandPL:OUTPUT
array([ 1, 10, 6, 24, 2, 23, 19, 12, 20, 22, 3, 9, 14, 13, 21, 7, 11, 15, 4, 16, 17, 8, 18, 5])OUTPUT
(24,) surveys_df[surveys_df["weight"] >= 0]-
OUTPUT
record_id month day year plot_id species_id sex hindfoot_length weight 13 14 7 16 1977 8 DM NaN NaN NaN 18 19 7 16 1977 4 PF NaN NaN NaN 33 34 7 17 1977 17 DM NaN NaN NaN 56 57 7 18 1977 22 DM NaN NaN NaN 76 77 8 19 1977 4 SS NaN NaN NaN ... ... ... ... ... ... ... ... ... ... 35527 35528 12 31 2002 13 US NaN NaN NaN 35543 35544 12 31 2002 15 US NaN NaN NaN 35544 35545 12 31 2002 15 AH NaN NaN NaN 35545 35546 12 31 2002 15 AH NaN NaN NaN 35548 35549 12 31 2002 5 NaN NaN NaN NaN [2511 rows x 9 columns]
When working through the solutions to the challenges above, you could introduce already that all these slice operations are actually based on a Boolean indexing operation (next section in the lesson). The filter provides for each record if it satisfies (True) or not (False). The slicing itself interprets the True/False of each record.
Using masks to identify a specific condition
A mask can be useful to locate where a particular
subset of values exist or don’t exist - for example, NaN, or “Not a
Number” values. To understand masks, we also need to understand
BOOLEAN objects in Python.
Boolean values include True or False. For
example,
When we ask Python whether x is greater than 5, it
returns False. This is Python’s way to say “No”. Indeed,
the value of x is 5, and 5 is not greater than 5.
To create a boolean mask:
- Set the True / False criteria
(e.g.
values > 5 = True) - Python will then assess each value in the object to determine whether the value meets the criteria (True) or not (False).
- Python creates an output object that is the same shape as the
original object, but with a
TrueorFalsevalue for each index location.
Let’s try this out. Let’s identify all locations in the survey data
that have null (missing or NaN) data values. We can use the
isnull method to do this. The isnull method
will compare each cell with a null value. If an element has a null
value, it will be assigned a value of True in the output
object.
A snippet of the output is below:
PYTHON
record_id month day year plot_id species_id sex hindfoot_length weight
0 False False False False False False False False True
1 False False False False False False False False True
2 False False False False False False False False True
3 False False False False False False False False True
4 False False False False False False False False True
[35549 rows x 9 columns]To select the rows where there are null values, we can use the mask as an index to subset our data as follows:
PYTHON
# To select just the rows with NaN values, we can use the 'any()' method
surveys_df[pd.isnull(surveys_df).any(axis=1)]Note that the weight column of our DataFrame contains
many null or NaN values. We will explore ways
of dealing with this in the next episode on Data Types and Formats.
We can run isnull on a particular column too. What does
the code below do?
PYTHON
# What does this do?
empty_weights = surveys_df[pd.isnull(surveys_df['weight'])]['weight']
print(empty_weights)Let’s take a minute to look at the statement above. We are using the
Boolean object pd.isnull(surveys_df['weight']) as an index
to surveys_df. We are asking Python to select rows that
have a NaN value of weight.
Challenge - Putting it all together
- Create a new DataFrame that only contains observations with sex values that are not female or male. Print the number of rows in this new DataFrame. Verify the result by comparing the number of rows in the new DataFrame with the number of rows in the surveys DataFrame where sex is null.
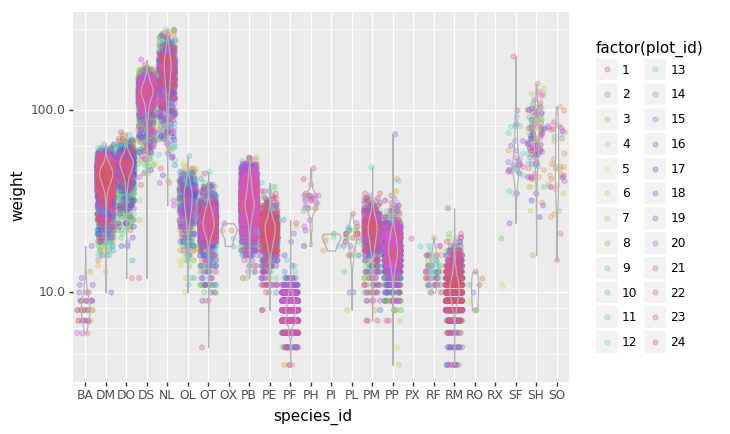
- Create a new DataFrame that contains only observations that are of sex male or female and where weight values are greater than 0. Create a stacked bar plot of average weight by plot with male vs female values stacked for each plot.
-
OUTPUT
2511OUTPUT
True PYTHON
# selection of the data with isin stack_selection = surveys_df[(surveys_df['sex'].isin(['M', 'F'])) & surveys_df["weight"] > 0.][["sex", "weight", "plot_id"]] # calculate the mean weight for each plot id and sex combination: stack_selection = stack_selection.groupby(["plot_id", "sex"]).mean().unstack() # and we can make a stacked bar plot from this: stack_selection.plot(kind='bar', stacked=True)
Referring to the challenge solution above, as we know the other values are all Nan values, we could also select all not null values:
PYTHON
stack_selection = surveys_df[(surveys_df['sex'].notnull()) &
surveys_df["weight"] > 0.][["sex", "weight", "plot_id"]]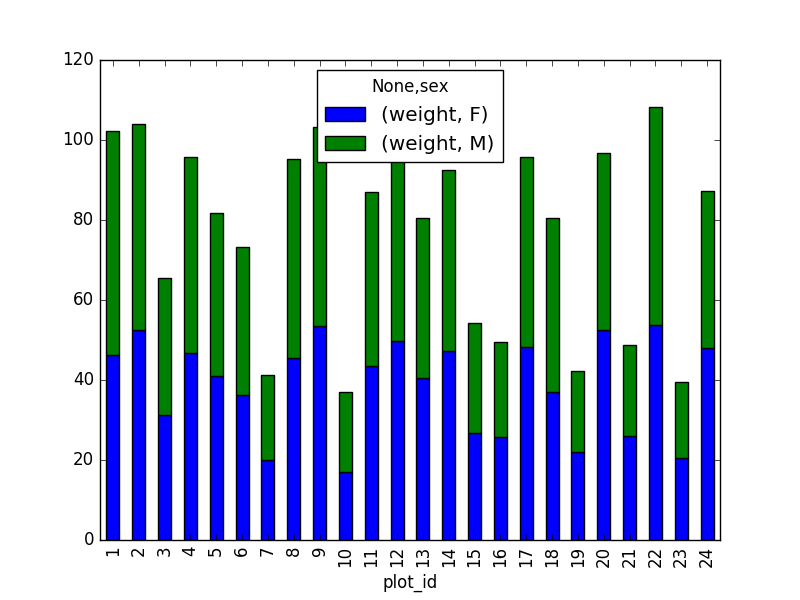
However, due to the unstack command, the legend header
contains two levels. In order to remove this, the column naming needs to
be simplified:
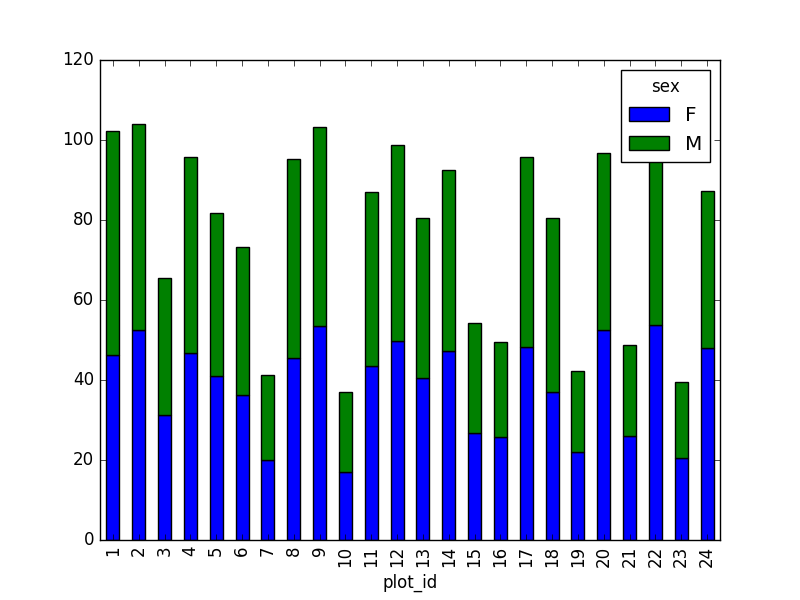
This is just a preview, more in next episode.
Key Points
- In Python, portions of data can be accessed using indices, slices, column headings, and condition-based subsetting.
- Python uses 0-based indexing, in which the first element in a list, tuple or any other data structure has an index of 0.
- Pandas enables common data exploration steps such as data indexing, slicing and conditional subsetting.
Content from Data Types and Formats
Last updated on 2023-06-05 | Edit this page
Estimated time: 45 minutes
Overview
Questions
- What types of data can be contained in a DataFrame?
- Why is the data type important?
Objectives
- Describe how information is stored in a pandas DataFrame.
- Define the two main types of data in pandas: text and numerics.
- Examine the structure of a DataFrame.
- Modify the format of values in a DataFrame.
- Describe how data types impact operations.
- Define, manipulate, and interconvert integers and floats in Python/pandas.
- Analyze datasets having missing/null values (NaN values).
- Write manipulated data to a file.
The format of individual columns and rows will impact analysis performed on a dataset read into a pandas DataFrame. For example, you can’t perform mathematical calculations on a string (text formatted data). This might seem obvious, however sometimes numeric values are read into pandas as strings. In this situation, when you then try to perform calculations on the string-formatted numeric data, you get an error.
In this lesson we will review ways to explore and better understand the structure and format of our data.
Types of Data
How information is stored in a DataFrame or a Python object affects what we can do with it and the outputs of calculations as well. There are two main types of data that we will explore in this lesson: numeric and text data types.
Numeric Data Types
Numeric data types include integers and floats. A floating point (known as a float) number has decimal points even if that decimal point value is 0. For example: 1.13, 2.0, 1234.345. If we have a column that contains both integers and floating point numbers, pandas will assign the entire column to the float data type so the decimal points are not lost.
An integer will never have a decimal point. Thus if
we wanted to store 1.13 as an integer it would be stored as 1.
Similarly, 1234.345 would be stored as 1234. You will often see the data
type Int64 in pandas which stands for 64 bit integer. The
64 refers to the memory allocated to store data in each cell which
effectively relates to how many digits it can store in each “cell”.
Allocating space ahead of time allows computers to optimize storage and
processing efficiency.
Text Data Type
The text data type is known as a string in Python, or
object in pandas. Strings can contain numbers and / or
characters. For example, a string might be a word, a sentence, or
several sentences. A pandas object might also be a plot name like
'plot1'. A string can also contain or consist of numbers.
For instance, '1234' could be stored as a string, as could
'10.23'. However strings that contain numbers can
not be used for mathematical operations!
pandas and base Python use slightly different names for data types. More on this is in the table below:
| Pandas Type | Native Python Type | Description |
|---|---|---|
| object | string | The most general dtype. Will be assigned to your column if column has mixed types (numbers and strings). |
| int64 | int | Numeric characters. 64 refers to the memory allocated to hold this character. |
| float64 | float | Numeric characters with decimals. If a column contains numbers and NaNs (see below), pandas will default to float64, in case your missing value has a decimal. |
| datetime64, timedelta[ns] | N/A (but see the datetime module in Python’s standard library) | Values meant to hold time data. Look into these for time series experiments. |
Checking the format of our data
Now that we’re armed with a basic understanding of numeric and text
data types, let’s explore the format of our survey data. We’ll be
working with the same surveys.csv dataset that we’ve used
in previous lessons.
PYTHON
# Make sure pandas is loaded
import pandas as pd
# Note that pd.read_csv is used because we imported pandas as pd
surveys_df = pd.read_csv("data/surveys.csv")Remember that we can check the type of an object like this:
OUTPUT
pandas.core.frame.DataFrameNext, let’s look at the structure of our surveys_df
data. In pandas, we can check the type of one column in a DataFrame
using the syntax dataframe_name['column_name'].dtype:
OUTPUT
dtype('O')A type ‘O’ just stands for “object” which in pandas is a string (text).
OUTPUT
dtype('int64')The type int64 tells us that pandas is storing each
value within this column as a 64 bit integer. We can use the
dataframe_name.dtypes command to view the data type for
each column in a DataFrame (all at once).
which returns:
PYTHON
record_id int64
month int64
day int64
year int64
plot_id int64
species_id object
sex object
hindfoot_length float64
weight float64
dtype: objectNote that most of the columns in our survey_df data are
of type int64. This means that they are 64 bit integers.
But the weight column is a floating point value which means
it contains decimals. The species_id and sex
columns are objects which means they contain strings.
Working With Integers and Floats
So we’ve learned that computers store numbers in one of two ways: as integers or as floating-point numbers (or floats). Integers are the numbers we usually count with. Floats have fractional parts (decimal places). Let’s next consider how the data type can impact mathematical operations on our data. Addition, subtraction, division and multiplication work on floats and integers as we’d expect.
OUTPUT
10OUTPUT
20If we divide one integer by another, we get a float. The result on Python 3 is different than in Python 2, where the result is an integer (integer division).
OUTPUT
0.5555555555555556OUTPUT
3.3333333333333335We can also convert a floating point number to an integer or an integer to floating point number. Notice that Python by default rounds down when it converts from floating point to integer.
OUTPUT
7OUTPUT
7.0Working With Our Survey Data
Getting back to our data, we can modify the format of values within
our data, if we want. For instance, we could convert the
record_id field to floating point values.
PYTHON
# Convert the record_id field from an integer to a float
surveys_df['record_id'] = surveys_df['record_id'].astype('float64')
surveys_df['record_id'].dtypeOUTPUT
dtype('float64')OUTPUT
0 2.0
1 3.0
2 2.0
3 7.0
4 3.0
...
35544 15.0
35545 15.0
35546 10.0
35547 7.0
35548 5.0ERROR
pandas.errors.IntCastingNaNError: Cannot convert non-finite values (NA or inf) to integerPandas cannot convert types from float to int if the column contains NaN values.
Missing Data Values - NaN
What happened in the last challenge activity? Notice that this raises
a casting error:
pandas.errors.IntCastingNaNError: Cannot convert non-finite values (NA or inf) to integer
(in older versions of pandas, this may be called a
ValueError instead). If we look at the weight
column in the surveys data we notice that there are NaN
(Not a Number)
values. NaN values are undefined values that cannot be
represented mathematically. pandas, for example, will read an empty cell
in a CSV or Excel sheet as NaN. NaNs have some desirable
properties: if we were to average the weight column without
replacing our NaNs, Python would know to skip over those cells.
OUTPUT
42.672428212991356Dealing with missing data values is always a challenge. It’s sometimes hard to know why values are missing - was it because of a data entry error? Or data that someone was unable to collect? Should the value be 0? We need to know how missing values are represented in the dataset in order to make good decisions. If we’re lucky, we have some metadata that will tell us more about how null values were handled.
For instance, in some disciplines, like Remote Sensing, missing data
values are often defined as -9999. Having a bunch of -9999 values in
your data could really alter numeric calculations. Often in
spreadsheets, cells are left empty where no data are available. pandas
will, by default, replace those missing values with NaN.
However, it is good practice to get in the habit of intentionally
marking cells that have no data with a no data value! That way there are
no questions in the future when you (or someone else) explores your
data.
Where Are the NaN’s?
Let’s explore the NaN values in our data a bit further.
Using the tools we learned in lesson 02, we can figure out how many rows
contain NaN values for weight. We can also
create a new subset from our data that only contains rows with
weight > 0 (i.e., select meaningful weight values):
PYTHON
len(surveys_df[surveys_df['weight'].isna()])
# How many rows have weight values?
len(surveys_df[surveys_df['weight'] > 0])We can replace all NaN values with zeroes using the
.fillna() method (after making a copy of the data so we
don’t lose our work):
However NaN and 0 yield different analysis
results. The mean value when NaN values are replaced with
0 is different from when NaN values are simply
thrown out or ignored.
OUTPUT
38.751976145601844We can fill NaN values with any value that we chose. The
code below fills all NaN values with a mean for all weight
values.
We could also chose to create a subset of our data, only keeping rows
that do not contain NaN values.
The point is to make conscious decisions about how to manage missing data. This is where we think about how our data will be used and how these values will impact the scientific conclusions made from the data.
pandas gives us all of the tools that we need to account for these issues. We just need to be cautious about how the decisions that we make impact scientific results.
Counting
Count the number of missing values per column.
The method .count() gives you the number of non-NaN
observations per column. Try looking to the .isna()
method.
Or, since we’ve been using the pd.isnull function so
far:
OUTPUT
record_id 0
month 0
day 0
year 0
plot_id 0
species_id 763
sex 2511
hindfoot_length 4111
weight 3266Note that isnull and isna are equivalent:
they behave identically.
Writing Out Data to CSV
We’ve learned about manipulating data to get desired outputs. But we’ve also discussed keeping data that has been manipulated separate from our raw data. Something we might be interested in doing is working with only the columns that have full data. First, let’s reload the data so we’re not mixing up all of our previous manipulations.
Next, let’s drop all the rows that contain missing values. We will
use the command dropna. By default, dropna
removes rows that contain missing data for even just one column.
If you now type df_na, you should observe that the
resulting DataFrame has 30676 rows and 9 columns, much smaller than the
35549 row original.
We can now use the to_csv command to export a DataFrame
in CSV format. Note that the code below will by default save the data
into the current working directory. We can save it to a different folder
by adding the foldername and a slash before the filename:
df.to_csv('foldername/out.csv'). We use
'index=False' so that pandas doesn’t include the index
number for each line.
We will use this data file later in the workshop. Check out your working directory to make sure the CSV wrote out properly, and that you can open it! If you want, try to bring it back into Python to make sure it imports properly.
If learners have trouble generating the output, or anything happens
with that, the folder sample_output
in this repository contains the file surveys_complete.csv
with the data they should generate.
Recap
What we’ve learned:
- How to explore the data types of columns within a DataFrame
- How to change the data type
- What NaN values are, how they might be represented, and what this means for your work
- How to replace NaN values, if desired
- How to use
to_csvto write manipulated data to a file.
Key Points
- pandas uses other names for data types than Python, for example:
objectfor textual data. - A column in a DataFrame can only have one data type.
- The data type in a DataFrame’s single column can be checked using
dtype. - Make conscious decisions about how to manage missing data.
- A DataFrame can be saved to a CSV file using the
to_csvfunction.
Content from Combining DataFrames with Pandas
Last updated on 2023-05-19 | Edit this page
Estimated time: 45 minutes
Overview
Questions
- Can I work with data from multiple sources?
- How can I combine data from different data sets?
Objectives
- Combine data from multiple files into a single DataFrame using merge and concat.
- Combine two DataFrames using a unique ID found in both DataFrames.
- Employ
to_csvto export a DataFrame in CSV format. - Join DataFrames using common fields (join keys).
In many “real world” situations, the data that we want to use come in
multiple files. We often need to combine these files into a single
DataFrame to analyze the data. The pandas package provides various
methods for combining DataFrames including merge and
concat.
To work through the examples below, we first need to load the species and surveys files into pandas DataFrames. In a Jupyter Notebook or iPython:
PYTHON
import pandas as pd
surveys_df = pd.read_csv("data/surveys.csv",
keep_default_na=False, na_values=[""])
surveys_dfOUTPUT
record_id month day year plot species sex hindfoot_length weight
0 1 7 16 1977 2 NA M 32 NaN
1 2 7 16 1977 3 NA M 33 NaN
2 3 7 16 1977 2 DM F 37 NaN
3 4 7 16 1977 7 DM M 36 NaN
4 5 7 16 1977 3 DM M 35 NaN
... ... ... ... ... ... ... ... ... ...
35544 35545 12 31 2002 15 AH NaN NaN NaN
35545 35546 12 31 2002 15 AH NaN NaN NaN
35546 35547 12 31 2002 10 RM F 15 14
35547 35548 12 31 2002 7 DO M 36 51
35548 35549 12 31 2002 5 NaN NaN NaN NaN
[35549 rows x 9 columns]PYTHON
species_df = pd.read_csv("data/species.csv",
keep_default_na=False, na_values=[""])
species_dfOUTPUT
species_id genus species taxa
0 AB Amphispiza bilineata Bird
1 AH Ammospermophilus harrisi Rodent
2 AS Ammodramus savannarum Bird
3 BA Baiomys taylori Rodent
4 CB Campylorhynchus brunneicapillus Bird
.. ... ... ... ...
49 UP Pipilo sp. Bird
50 UR Rodent sp. Rodent
51 US Sparrow sp. Bird
52 ZL Zonotrichia leucophrys Bird
53 ZM Zenaida macroura Bird
[54 rows x 4 columns]Take note that the read_csv method we used can take some
additional options which we didn’t use previously. Many functions in
Python have a set of options that can be set by the user if needed. In
this case, we have told pandas to assign empty values in our CSV to
NaN with the parameters keep_default_na=False
and na_values=[""]. We have explicitly requested to change
empty values in the CSV to NaN, this is however also the default
behaviour of read_csv. More
about all of the read_csv options here and their
defaults.
Concatenating DataFrames
We can use the concat function in pandas to append
either columns or rows from one DataFrame to another. Let’s grab two
subsets of our data to see how this works.
PYTHON
# Read in first 10 lines of surveys table
survey_sub = surveys_df.head(10)
# Grab the last 10 rows
survey_sub_last10 = surveys_df.tail(10)
# Reset the index values to the second dataframe appends properly
survey_sub_last10 = survey_sub_last10.reset_index(drop=True)
# drop=True option avoids adding new index column with old index valuesWhen we concatenate DataFrames, we need to specify the axis.
axis=0 tells pandas to stack the second DataFrame UNDER the
first one. It will automatically detect whether the column names are the
same and will stack accordingly. axis=1 will stack the
columns in the second DataFrame to the RIGHT of the first DataFrame. To
stack the data vertically, we need to make sure we have the same columns
and associated column format in both datasets. When we stack
horizontally, we want to make sure what we are doing makes sense
(i.e. the data are related in some way).
PYTHON
# Stack the DataFrames on top of each other
vertical_stack = pd.concat([survey_sub, survey_sub_last10], axis=0)
# Place the DataFrames side by side
horizontal_stack = pd.concat([survey_sub, survey_sub_last10], axis=1)Row Index Values and Concat
Have a look at the vertical_stack DataFrame. Notice
anything unusual? The row indexes for the two DataFrames
survey_sub and survey_sub_last10 have been
repeated. We can reindex the new DataFrame using the
reset_index() method.
Writing Out Data to CSV
We can use the to_csv command to do export a DataFrame
in CSV format. Note that the code below will by default save the data
into the current working directory. We can save it to a different folder
by adding the foldername and a slash to the file
vertical_stack.to_csv('foldername/out.csv'). We use the
index=False so that pandas doesn’t include the index number
for each line.
Check out your working directory to make sure the CSV wrote out properly, and that you can open it! If you want, try to bring it back into pandas to make sure it imports properly.
PYTHON
# For kicks read our output back into Python and make sure all looks good
new_output = pd.read_csv('data/out.csv', keep_default_na=False, na_values=[""])Challenge - Combine Data
In the data folder, there is another folder called
yearly_files that contains survey data broken down into
individual files by year. Read the data from two of these files,
surveys2001.csv and surveys2002.csv, into
pandas and combine the files to make one new DataFrame. Create a plot of
average plot weight by year grouped by sex. Export your results as a CSV
and make sure it reads back into pandas properly.
PYTHON
# read the files:
survey2001 = pd.read_csv("data/yearly_files/surveys2001.csv")
survey2002 = pd.read_csv("data/yearly_files/surveys2002.csv")
# concatenate
survey_all = pd.concat([survey2001, survey2002], axis=0)
# get the weight for each year, grouped by sex:
weight_year = survey_all.groupby(['year', 'sex']).mean()["wgt"].unstack()
# plot:
weight_year.plot(kind="bar")
plt.tight_layout() # tip: use this to improve the plot layout.
# Try running the code without this line to see
# what difference applying plt.tight_layout() makes.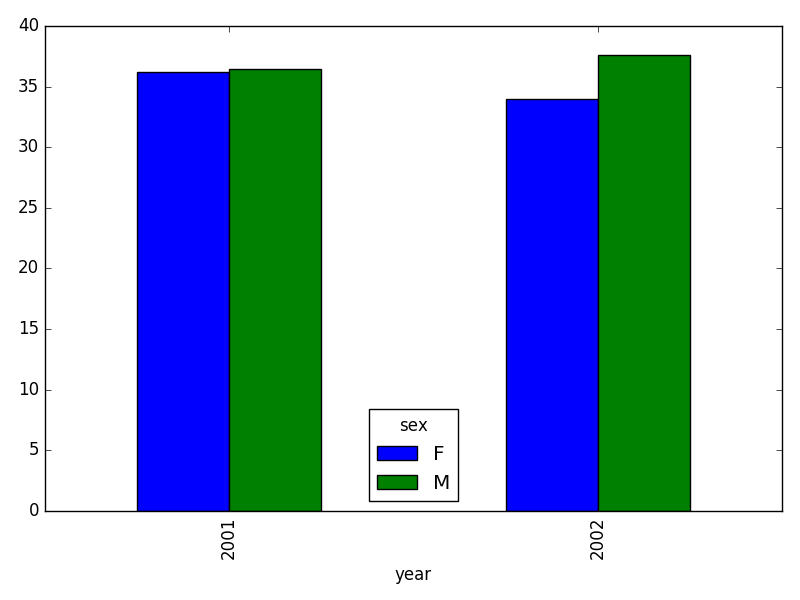
Joining DataFrames
When we concatenated our DataFrames, we simply added them to each other - stacking them either vertically or side by side. Another way to combine DataFrames is to use columns in each dataset that contain common values (a common unique identifier). Combining DataFrames using a common field is called “joining”. The columns containing the common values are called “join key(s)”. Joining DataFrames in this way is often useful when one DataFrame is a “lookup table” containing additional data that we want to include in the other.
NOTE: This process of joining tables is similar to what we do with tables in an SQL database.
For example, the species.csv file that we’ve been
working with is a lookup table. This table contains the genus, species
and taxa code for 55 species. The species code is unique for each line.
These species are identified in our survey data as well using the unique
species code. Rather than adding three more columns for the genus,
species and taxa to each of the 35,549 line survey
DataFrame, we can maintain the shorter table with the species
information. When we want to access that information, we can create a
query that joins the additional columns of information to the
survey DataFrame.
Storing data in this way has many benefits.
- It ensures consistency in the spelling of species attributes (genus, species and taxa) given each species is only entered once. Imagine the possibilities for spelling errors when entering the genus and species thousands of times!
- It also makes it easy for us to make changes to the species information once without having to find each instance of it in the larger survey data.
- It optimizes the size of our data.
Joining Two DataFrames
To better understand joins, let’s grab the first 10 lines of our data
as a subset to work with. We’ll use the .head() method to
do this. We’ll also read in a subset of the species table.
PYTHON
# Read in first 10 lines of surveys table
survey_sub = surveys_df.head(10)
# Import a small subset of the species data designed for this part of the lesson.
# It is stored in the data folder.
species_sub = pd.read_csv('data/speciesSubset.csv', keep_default_na=False, na_values=[""])In this example, species_sub is the lookup table
containing genus, species, and taxa names that we want to join with the
data in survey_sub to produce a new DataFrame that contains
all of the columns from both species_df and
survey_df.
Identifying join keys
To identify appropriate join keys we first need to know which field(s) are shared between the files (DataFrames). We might inspect both DataFrames to identify these columns. If we are lucky, both DataFrames will have columns with the same name that also contain the same data. If we are less lucky, we need to identify a (differently-named) column in each DataFrame that contains the same information.
OUTPUT
Index([u'species_id', u'genus', u'species', u'taxa'], dtype='object')OUTPUT
Index([u'record_id', u'month', u'day', u'year', u'plot_id', u'species_id',
u'sex', u'hindfoot_length', u'weight'], dtype='object')In our example, the join key is the column containing the two-letter
species identifier, which is called species_id.
Now that we know the fields with the common species ID attributes in each DataFrame, we are almost ready to join our data. However, since there are different types of joins, we also need to decide which type of join makes sense for our analysis.
Inner joins
The most common type of join is called an inner join. An inner join combines two DataFrames based on a join key and returns a new DataFrame that contains only those rows that have matching values in both of the original DataFrames. An example of an inner join, adapted from Jeff Atwood’s blogpost about SQL joins is below:
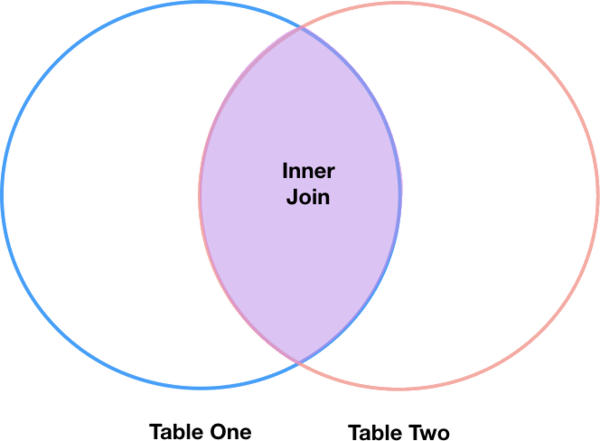
The pandas function for performing joins is called merge
and an Inner join is the default option:
PYTHON
merged_inner = pd.merge(left=survey_sub, right=species_sub, left_on='species_id', right_on='species_id')In this case, species_id is the only column name in both
DataFrames, so if we skipped the left_on and
right_on arguments, pandas would guess that we
wanted to use that column to join. However, it is usually better to be
explicit.
So what is the size of the output data?
OUTPUT
(8, 12)OUTPUT
record_id month day year plot_id species_id sex hindfoot_length \
0 1 7 16 1977 2 NL M 32
1 2 7 16 1977 3 NL M 33
2 3 7 16 1977 2 DM F 37
3 4 7 16 1977 7 DM M 36
4 5 7 16 1977 3 DM M 35
5 8 7 16 1977 1 DM M 37
6 9 7 16 1977 1 DM F 34
7 7 7 16 1977 2 PE F NaN
weight genus species taxa
0 NaN Neotoma albigula Rodent
1 NaN Neotoma albigula Rodent
2 NaN Dipodomys merriami Rodent
3 NaN Dipodomys merriami Rodent
4 NaN Dipodomys merriami Rodent
5 NaN Dipodomys merriami Rodent
6 NaN Dipodomys merriami Rodent
7 NaN Peromyscus eremicus RodentThe result of an inner join of survey_sub and
species_sub is a new DataFrame that contains the combined
set of columns from survey_sub and
species_sub. It only contains rows that have
two-letter species codes that are the same in both the
survey_sub and species_sub DataFrames. In
other words, if a row in survey_sub has a value of
species_id that does not appear in the
species_id column of species, it will not be
included in the DataFrame returned by an inner join. Similarly, if a row
in species_sub has a value of species_id that
does not appear in the species_id column of
survey_sub, that row will not be included in the DataFrame
returned by an inner join.
The two DataFrames that we want to join are passed to the
merge function using the left and
right argument. The left_on='species_id'
argument tells merge to use the species_id
column as the join key from survey_sub (the
left DataFrame). Similarly , the
right_on='species_id' argument tells merge to
use the species_id column as the join key from
species_sub (the right DataFrame). For inner
joins, the order of the left and right
arguments does not matter.
The result merged_inner DataFrame contains all of the
columns from survey_sub (record_id,
month, day, etc.) as well as all the columns
from species_sub (species_id,
genus, species, and taxa).
Notice that merged_inner has fewer rows than
survey_sub. This is an indication that there were rows in
surveys_df with value(s) for species_id that
do not exist as value(s) for species_id in
species_df.
Left joins
What if we want to add information from species_sub to
survey_sub without losing any of the information from
survey_sub? In this case, we use a different type of join
called a “left outer join”, or a “left join”.
Like an inner join, a left join uses join keys to combine two
DataFrames. Unlike an inner join, a left join will return all
of the rows from the left DataFrame, even those rows whose
join key(s) do not have values in the right DataFrame. Rows
in the left DataFrame that are missing values for the join
key(s) in the right DataFrame will simply have null (i.e.,
NaN or None) values for those columns in the resulting joined
DataFrame.
Note: a left join will still discard rows from the right
DataFrame that do not have values for the join key(s) in the
left DataFrame.
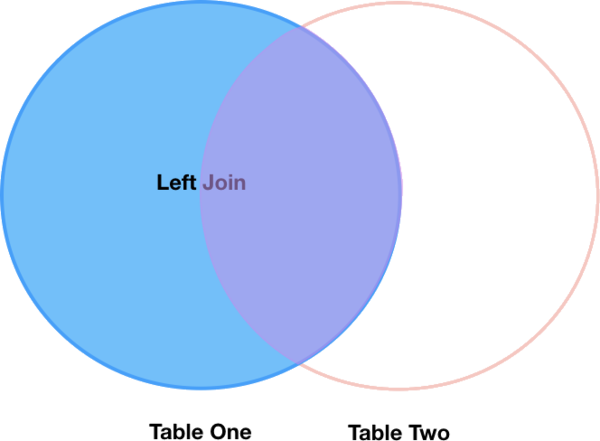

A left join is performed in pandas by calling the same
merge function used for inner join, but using the
how='left' argument:
PYTHON
merged_left = pd.merge(left=survey_sub, right=species_sub, how='left', left_on='species_id', right_on='species_id')
merged_leftOUTPUT
record_id month day year plot_id species_id sex hindfoot_length \
0 1 7 16 1977 2 NL M 32
1 2 7 16 1977 3 NL M 33
2 3 7 16 1977 2 DM F 37
3 4 7 16 1977 7 DM M 36
4 5 7 16 1977 3 DM M 35
5 6 7 16 1977 1 PF M 14
6 7 7 16 1977 2 PE F NaN
7 8 7 16 1977 1 DM M 37
8 9 7 16 1977 1 DM F 34
9 10 7 16 1977 6 PF F 20
weight genus species taxa
0 NaN Neotoma albigula Rodent
1 NaN Neotoma albigula Rodent
2 NaN Dipodomys merriami Rodent
3 NaN Dipodomys merriami Rodent
4 NaN Dipodomys merriami Rodent
5 NaN NaN NaN NaN
6 NaN Peromyscus eremicus Rodent
7 NaN Dipodomys merriami Rodent
8 NaN Dipodomys merriami Rodent
9 NaN NaN NaN NaNThe result DataFrame from a left join (merged_left)
looks very much like the result DataFrame from an inner join
(merged_inner) in terms of the columns it contains.
However, unlike merged_inner, merged_left
contains the same number of rows as the original
survey_sub DataFrame. When we inspect
merged_left, we find there are rows where the information
that should have come from species_sub (i.e.,
species_id, genus, and taxa) is
missing (they contain NaN values):
OUTPUT
record_id month day year plot_id species_id sex hindfoot_length \
5 6 7 16 1977 1 PF M 14
9 10 7 16 1977 6 PF F 20
weight genus species taxa
5 NaN NaN NaN NaN
9 NaN NaN NaN NaNThese rows are the ones where the value of species_id
from survey_sub (in this case, PF) does not
occur in species_sub.
Other join types
The pandas merge function supports two other join
types:
- Right (outer) join: Invoked by passing
how='right'as an argument. Similar to a left join, except all rows from therightDataFrame are kept, while rows from theleftDataFrame without matching join key(s) values are discarded. - Full (outer) join: Invoked by passing
how='outer'as an argument. This join type returns the all pairwise combinations of rows from both DataFrames; i.e., the result DataFrame willNaNwhere data is missing in one of the dataframes. This join type is very rarely used.
Final Challenges
Challenge - Distributions
Create a new DataFrame by joining the contents of the
surveys.csv and species.csv tables. Then
calculate and plot the distribution of:
- taxa by plot
- taxa by sex by plot
- taxa per plot (number of species of each taxa per plot):
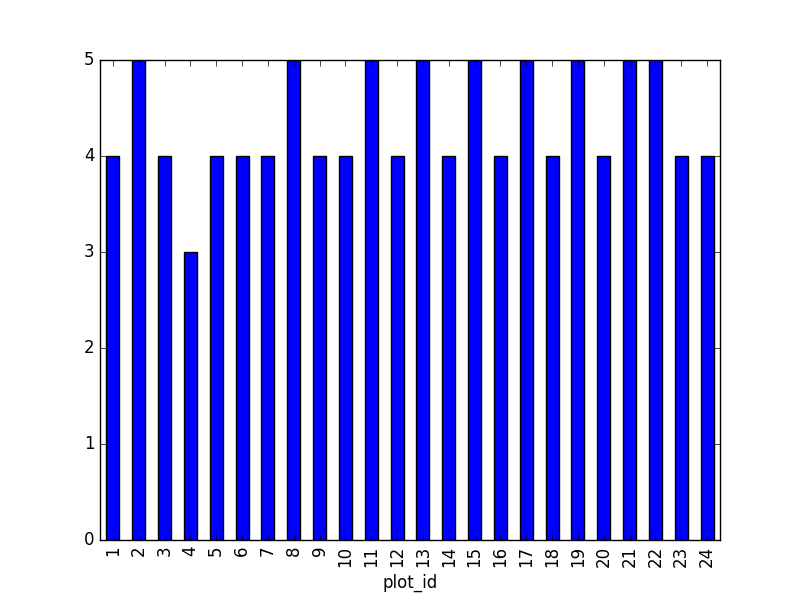
Suggestion: It is also possible to plot the number of individuals for each taxa in each plot (stacked bar chart):
PYTHON
merged_left.groupby(["plot_id", "taxa"]).count()["record_id"].unstack().plot(kind='bar', stacked=True)
plt.legend(loc='upper center', ncol=3, bbox_to_anchor=(0.5, 1.05)) # stop the legend from overlapping with the bar plot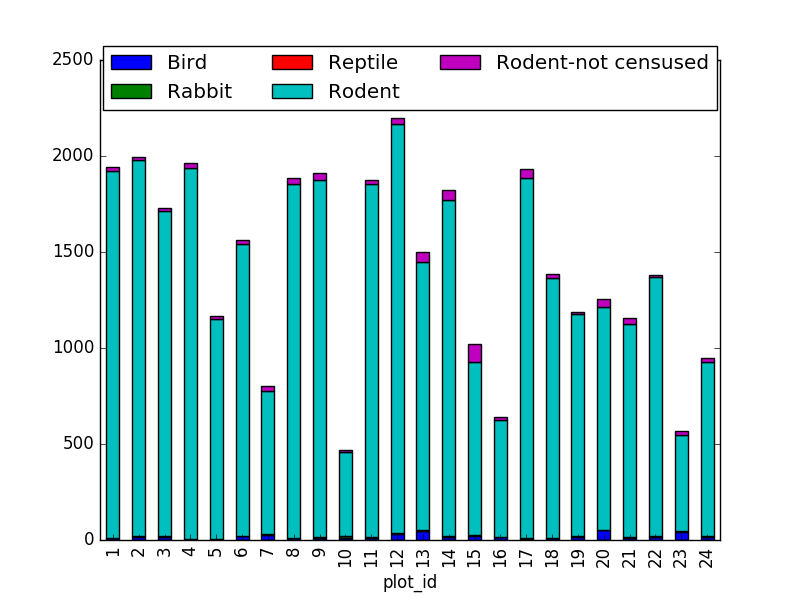
- taxa by sex by plot: Providing the Nan values with the M|F values (can also already be changed to ‘x’):
PYTHON
merged_left.loc[merged_left["sex"].isnull(), "sex"] = 'M|F'
ntaxa_sex_site= merged_left.groupby(["plot_id", "sex"])["taxa"].nunique().reset_index(level=1)
ntaxa_sex_site = ntaxa_sex_site.pivot_table(values="taxa", columns="sex", index=ntaxa_sex_site.index)
ntaxa_sex_site.plot(kind="bar", legend=False, stacked=True)
plt.legend(loc='upper center', ncol=3, bbox_to_anchor=(0.5, 1.08),
fontsize='small', frameon=False)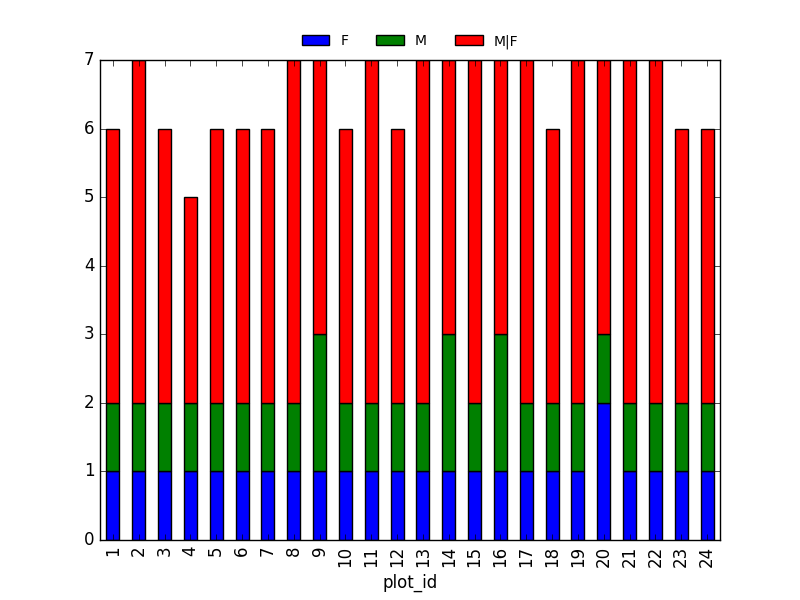
The number of individuals for each taxa in each plot per sex can be derived as well.
PYTHON
sex_taxa_site = merged_left.groupby(["plot_id", "taxa", "sex"]).count()['record_id']
sex_taxa_site.unstack(level=[1, 2]).plot(kind='bar', logy=True)
plt.legend(loc='upper center', ncol=3, bbox_to_anchor=(0.5, 1.15),
fontsize='small', frameon=False)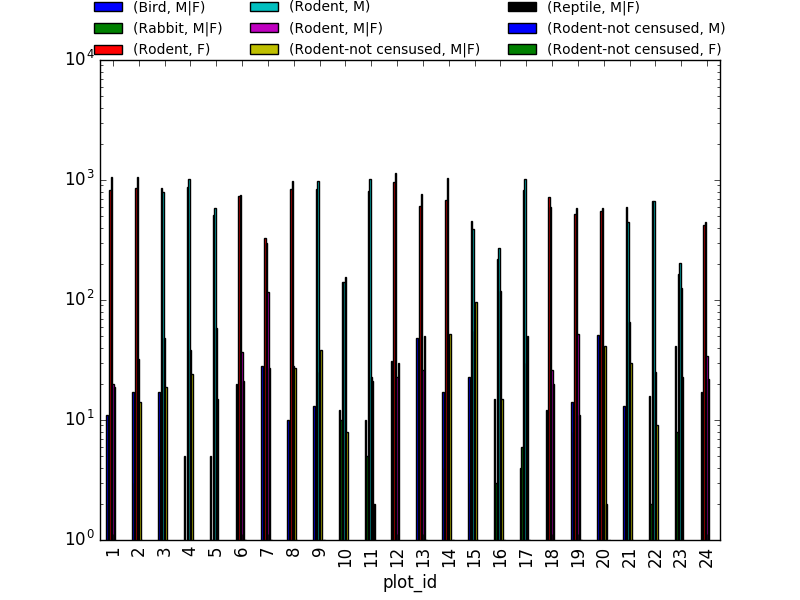
This is not really the best plot choice, e.g. it is not easily
readable. A first option to make this better, is to make facets.
However, pandas/matplotlib do not provide this by default. Just as a
pure matplotlib example (M|F if for not-defined sex
records):
PYTHON
fig, axs = plt.subplots(3, 1)
for sex, ax in zip(["M", "F", "M|F"], axs):
sex_taxa_site[sex_taxa_site["sex"] == sex].plot(kind='bar', ax=ax, legend=False)
ax.set_ylabel(sex)
if not ax.is_last_row():
ax.set_xticks([])
ax.set_xlabel("")
axs[0].legend(loc='upper center', ncol=5, bbox_to_anchor=(0.5, 1.3),
fontsize='small', frameon=False)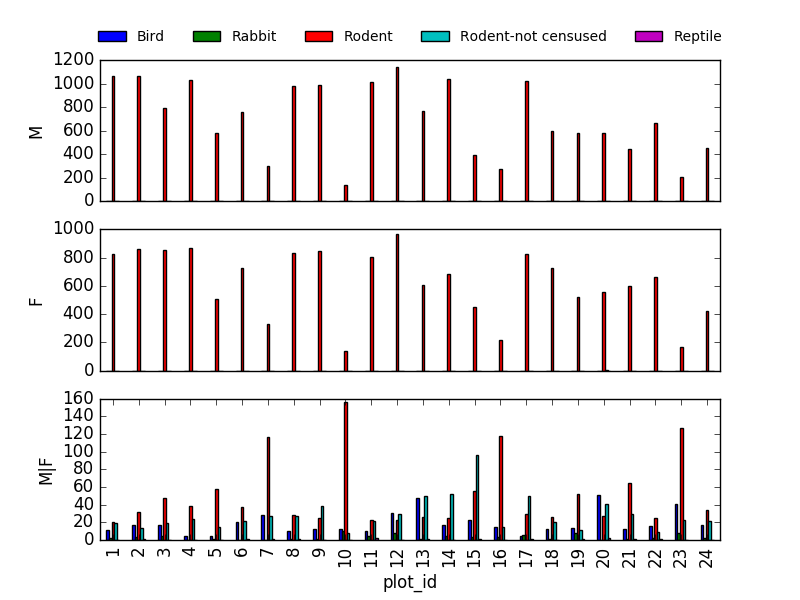
However, it would be better to link to Seaborn and Altair for this kind of multivariate visualisation.
Challenge - Diversity Index
In the data folder, there is a
plots.csvfile that contains information about the type associated with each plot. Use that data to summarize the number of plots by plot type.Calculate a diversity index of your choice for control vs rodent exclosure plots. The index should consider both species abundance and number of species. You might choose to use the simple biodiversity index described here which calculates diversity as: the number of species in the plot / the total number of individuals in the plot = Biodiversity index.
PYTHON
merged_site_type = pd.merge(merged_left, plot_info, on='plot_id')
# For each plot, get the number of species for each plot
nspecies_site = merged_site_type.groupby(["plot_id"])["species"].nunique().rename("nspecies")
# For each plot, get the number of individuals
nindividuals_site = merged_site_type.groupby(["plot_id"]).count()['record_id'].rename("nindiv")
# combine the two series
diversity_index = pd.concat([nspecies_site, nindividuals_site], axis=1)
# calculate the diversity index
diversity_index['diversity'] = diversity_index['nspecies']/diversity_index['nindiv']Making a bar chart from this diversity index:
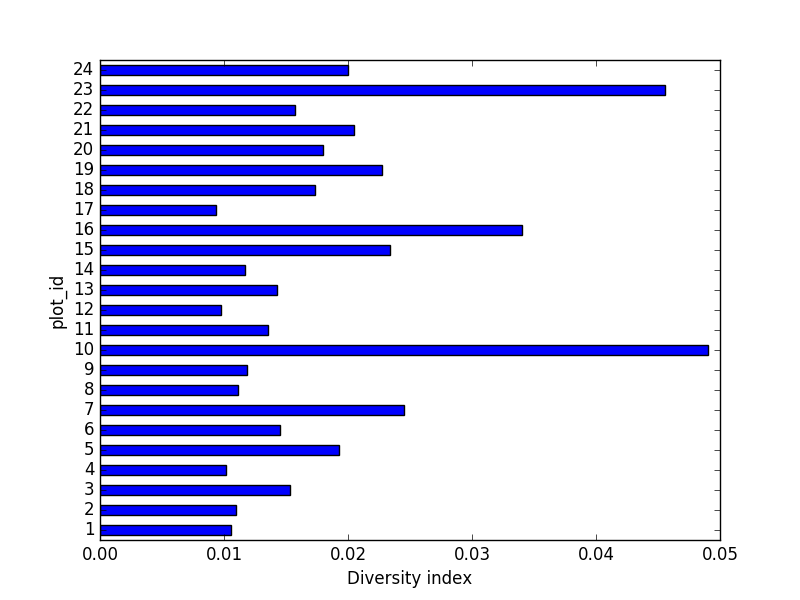
Key Points
- Pandas’
mergeandconcatcan be used to combine subsets of a DataFrame, or even data from different files. -
joinfunction combines DataFrames based on index or column. - Joining two DataFrames can be done in multiple ways (left, right, and inner) depending on what data must be in the final DataFrame.
-
to_csvcan be used to write out DataFrames in CSV format.
Content from Data Workflows and Automation
Last updated on 2023-05-18 | Edit this page
Estimated time: 90 minutes
Overview
Questions
- Can I automate operations in Python?
- What are functions and why should I use them?
Objectives
- Describe why for loops are used in Python.
- Employ for loops to automate data analysis.
- Write unique filenames in Python.
- Build reusable code in Python.
- Write functions using conditional statements (if, then, else).
So far, we’ve used Python and the pandas library to explore and manipulate individual datasets by hand, much like we would do in a spreadsheet. The beauty of using a programming language like Python, though, comes from the ability to automate data processing through the use of loops and functions.
For loops
Loops allow us to repeat a workflow (or series of actions) a given number of times or while some condition is true. We would use a loop to automatically process data that’s stored in multiple files (daily values with one file per year, for example). Loops lighten our work load by performing repeated tasks without our direct involvement and make it less likely that we’ll introduce errors by making mistakes while processing each file by hand.
Let’s write a simple for loop that simulates what a kid might see during a visit to the zoo:
OUTPUT
['lion', 'tiger', 'crocodile', 'vulture', 'hippo']OUTPUT
lion
tiger
crocodile
vulture
hippoThe line defining the loop must start with for and end
with a colon, and the body of the loop must be indented.
In this example, creature is the loop variable that
takes the value of the next entry in animals every time the
loop goes around. We can call the loop variable anything we like. After
the loop finishes, the loop variable will still exist and will have the
value of the last entry in the collection:
OUTPUT
OUTPUT
The loop variable is now: hippoWe are not asking Python to print the value of the loop variable
anymore, but the for loop still runs and the value of
creature changes on each pass through the loop. The
statement pass in the body of the loop means “do
nothing”.
Challenge - Loops
What happens if we don’t include the
passstatement?Rewrite the loop so that the animals are separated by commas, not new lines (Hint: You can concatenate strings using a plus sign. For example,
print(string1 + string2)outputs ‘string1string2’).
-
ERROR
IndentationError: expected an indented block -
Using the
endargument toprint:OUTPUT
lion,tiger,crocodile,vulture,hippo,This puts a comma on the end of the list, which is not ideal. To avoid this, we need to use an altogether different approach: string objects in Python have a
joinmethod, which can be used to concatenate items in a list with the string in between, e.g.OUTPUT
'lion, tiger, crocodile, vulture, hippo'
Automating data processing using For Loops
The file we’ve been using so far, surveys.csv, contains
25 years of data and is very large. We would like to separate the data
for each year into a separate file.
Let’s start by making a new directory inside the folder
data to store all of these files using the module
os:
The command os.mkdir is equivalent to mkdir
in the shell. Just so we are sure, we can check that the new directory
was created within the data folder:
OUTPUT
['plots.csv',
'portal_mammals.sqlite',
'species.csv',
'survey2001.csv',
'survey2002.csv',
'surveys.csv',
'surveys2002_temp.csv',
'yearly_files']The command os.listdir is equivalent to ls
in the shell.
In previous lessons, we saw how to use the library pandas to load the species data into memory as a DataFrame, how to select a subset of the data using some criteria, and how to write the DataFrame into a CSV file. Let’s write a script that performs those three steps in sequence for the year 2002:
PYTHON
import pandas as pd
# Load the data into a DataFrame
surveys_df = pd.read_csv('data/surveys.csv')
# Select only data for the year 2002
surveys2002 = surveys_df[surveys_df.year == 2002]
# Write the new DataFrame to a CSV file
surveys2002.to_csv('data/yearly_files/surveys2002.csv')To create yearly data files, we could repeat the last two commands over and over, once for each year of data. Repeating code is neither elegant nor practical, and is very likely to introduce errors into your code. We want to turn what we’ve just written into a loop that repeats the last two commands for every year in the dataset.
Let’s start by writing a loop that prints the names of the files we want to create - the dataset we are using covers 1977 through 2002, and we’ll create a separate file for each of those years. Listing the filenames is a good way to confirm that the loop is behaving as we expect.
We have seen that we can loop over a list of items, so we need a list of years to loop over. We can get the years in our DataFrame with:
OUTPUT
0 1977
1 1977
2 1977
3 1977
...
35545 2002
35546 2002
35547 2002
35548 2002but we want only unique years, which we can get using the
unique method which we have already seen.
OUTPUT
array([1977, 1978, 1979, 1980, 1981, 1982, 1983, 1984, 1985, 1986, 1987,
1988, 1989, 1990, 1991, 1992, 1993, 1994, 1995, 1996, 1997, 1998,
1999, 2000, 2001, 2002], dtype=int64)Putting this into our for loop we get
PYTHON
for year in surveys_df['year'].unique():
filename='data/yearly_files/surveys' + str(year) + '.csv'
print(filename)OUTPUT
data/yearly_files/surveys1977.csv
data/yearly_files/surveys1978.csv
data/yearly_files/surveys1979.csv
data/yearly_files/surveys1980.csv
data/yearly_files/surveys1981.csv
data/yearly_files/surveys1982.csv
data/yearly_files/surveys1983.csv
data/yearly_files/surveys1984.csv
data/yearly_files/surveys1985.csv
data/yearly_files/surveys1986.csv
data/yearly_files/surveys1987.csv
data/yearly_files/surveys1988.csv
data/yearly_files/surveys1989.csv
data/yearly_files/surveys1990.csv
data/yearly_files/surveys1991.csv
data/yearly_files/surveys1992.csv
data/yearly_files/surveys1993.csv
data/yearly_files/surveys1994.csv
data/yearly_files/surveys1995.csv
data/yearly_files/surveys1996.csv
data/yearly_files/surveys1997.csv
data/yearly_files/surveys1998.csv
data/yearly_files/surveys1999.csv
data/yearly_files/surveys2000.csv
data/yearly_files/surveys2001.csv
data/yearly_files/surveys2002.csvWe can now add the rest of the steps we need to create separate text files:
PYTHON
# Load the data into a DataFrame
surveys_df = pd.read_csv('data/surveys.csv')
for year in surveys_df['year'].unique():
# Select data for the year
surveys_year = surveys_df[surveys_df.year == year]
# Write the new DataFrame to a CSV file
filename = 'data/yearly_files/surveys' + str(year) + '.csv'
surveys_year.to_csv(filename)Look inside the yearly_files directory and check a
couple of the files you just created to confirm that everything worked
as expected.
Writing Unique File Names
Notice that the code above created a unique filename for each year.
Let’s break down the parts of this name:
- The first part is some text that specifies the directory to store
our data file in (data/yearly_files/) and the first part of the file
name (surveys):
'data/yearly_files/surveys' - We can concatenate this with the value of a variable, in this case
yearby using the plus+sign and the variable we want to add to the file name:+ str(year) - Then we add the file extension as another text string:
+ '.csv'
Notice that we use single quotes to add text strings. The variable is
not surrounded by quotes. This code produces the string
data/yearly_files/surveys2002.csv which contains the path
to the new filename AND the file name itself.
Challenge - Modifying loops
Some of the surveys you saved are missing data (they have null values that show up as NaN - Not A Number - in the DataFrames and do not show up in the text files). Modify the for loop so that the entries with null values are not included in the yearly files.
Let’s say you only want to look at data from a given multiple of years. How would you modify your loop in order to generate a data file for only every 5th year, starting from 1977?
Instead of splitting out the data by years, a colleague wants to do analyses each species separately. How would you write a unique CSV file for each species?
-
You could just make a list manually, however, why not check the first and last year making use of the code itself?
Building reusable and modular code with functions
Suppose that separating large data files into individual yearly files is a task that we frequently have to perform. We could write a for loop like the one above every time we needed to do it but that would be time consuming and error prone. A more elegant solution would be to create a reusable tool that performs this task with minimum input from the user. To do this, we are going to turn the code we’ve already written into a function.
Functions are reusable, self-contained pieces of code that are called with a single command. They can be designed to accept arguments as input and return values, but they don’t need to do either. Variables declared inside functions only exist while the function is running and if a variable within the function (a local variable) has the same name as a variable somewhere else in the code, the local variable hides but doesn’t overwrite the other.
Every method used in Python (for example, print) is a
function, and the libraries we import (say, pandas) are a
collection of functions. We will only use functions that are housed
within the same code that uses them, but we can also write functions
that can be used by different programs.
Functions are declared following this general structure:
PYTHON
def this_is_the_function_name(input_argument1, input_argument2):
# The body of the function is indented
# This function prints the two arguments to screen
print('The function arguments are:', input_argument1, input_argument2, '(this is done inside the function!)')
# And returns their product
return input_argument1 * input_argument2The function declaration starts with the word def,
followed by the function name and any arguments in parenthesis, and ends
in a colon. The body of the function is indented just like loops are. If
the function returns something when it is called, it includes a return
statement at the end.
This is how we call the function:
OUTPUT
The function arguments are: 2 5 (this is done inside the function!)OUTPUT
Their product is: 10 (this is done outside the function!)When teaching the challenge below, demonstrating the results in a debugging environment can make function scope more clear.
Challenge - Functions
- Change the values of the arguments provided to the function and check its output
- Try calling the function by giving it the wrong number of arguments
(not 2) or not assigning the function call to a variable (no
product_of_inputs =) - Declare a variable inside the function and test to see where it exists (Hint: can you print it from outside the function?)
- Explore what happens when a variable both inside and outside the function have the same name. What happens to the global variable when you change the value of the local variable?
-
For example, using an integer and a string:
OUTPUT
The function arguments are: 3 clap (this is done inside the function!) clapclapclap -
For example, providing a third integer argument:
ERROR
TypeError: this_is_the_function_name() takes 2 positional arguments but 3 were givenand proving the correct number of arguments but not capturing the return value in a variable:
OUTPUT
The function arguments are: 23 108 (this is done inside the function!) 2484 -
An example looking at variables defined outside and inside the function. Note that the explanatory comments have been removed from the function definition.
PYTHON
var_defined_outside = 'outside' def this_is_the_function_name(input_argument1, input_argument2): var_defined_inside = 'inside' print('The function arguments are:', input_argument1, input_argument2, '(this is done inside the function!)') print('This variable was created ' + var_defined_outside + ' the function') return input_argument1 * input_argument2 this_is_the_function_name(3.3, 7.9) print('This variable was created' + var_defined_inside + ' the function')OUTPUT
The function arguments are: 3.3 7.9 (this is done inside the function!) This variable was created outside the function 26.07ERROR
NameError: name 'var_defined_inside' is not definedWe can see from these results that variables defined outside the function are available for use inside them, while variables defined inside a function are not available outside them.
-
PYTHON
shared_variable_name = 'who would I be' def this_is_the_function_name(input_argument1, input_argument2): shared_variable_name = 'without you' print('The function arguments are:', input_argument1, input_argument2, '(this is done inside the function!)') return input_argument1 * input_argument2 print(shared_variable_name) this_is_the_function_name(2, 3) # does calling the function change the variable's value? print(shared_variable_name)OUTPUT
who would I be The function arguments are: 2 3 (this is done inside the function!) 6 who would I beAnd what about if we modify the variable inside the function?
PYTHON
def this_is_the_function_name(input_argument1, input_argument2): shared_variable_name = 'without you' shared_variable_name = shared_variable_name + ', without them?' print(shared_variable_name) print('The function arguments are:', input_argument1, input_argument2, '(this is done inside the function!)') return input_argument1 * input_argument2 this_is_the_function_name(2, 3) print(shared_variable_name)OUTPUT
without you, without them? The function arguments are: 2 3 (this is done inside the function!) 6 who would I be
We can now turn our code for saving yearly data files into a function. There are many different “chunks” of this code that we can turn into functions, and we can even create functions that call other functions inside them. Let’s first write a function that separates data for just one year and saves that data to a file:
PYTHON
def one_year_csv_writer(this_year, all_data):
"""
Writes a csv file for data from a given year.
this_year -- year for which data is extracted
all_data -- DataFrame with multi-year data
"""
# Select data for the year
surveys_year = all_data[all_data.year == this_year]
# Write the new DataFrame to a csv file
filename = 'data/yearly_files/function_surveys' + str(this_year) + '.csv'
surveys_year.to_csv(filename)The text between the two sets of triple double quotes is called a
docstring and contains the documentation for the function. It does
nothing when the function is running and is therefore not necessary, but
it is good practice to include docstrings as a reminder of what the code
does. Docstrings in functions also become part of their ‘official’
documentation, and we can see them by typing
help(function_name):
OUTPUT
Help on function one_year_csv_writer in module __main__:
one_year_csv_writer(this_year, all_data)
Writes a csv file for data from a given year.
this_year -- year for which data is extracted
all_data -- DataFrame with multi-year dataOr, when working in the Jupyter environment, adding a ?
(question mark) after the function name:
We changed the root of the name of the CSV file so we can distinguish
it from the one we wrote before. Check the yearly_files
directory for the file. Did it do what you expect?
What we really want to do, though, is create files for multiple years
without having to request them one by one. Let’s write another function
that replaces the entire for loop by looping through a
sequence of years and repeatedly calling the function we just wrote,
one_year_csv_writer:
PYTHON
def yearly_data_csv_writer(start_year, end_year, all_data):
"""
Writes separate CSV files for each year of data.
start_year -- the first year of data we want
end_year -- the last year of data we want
all_data -- DataFrame with multi-year data
"""
# "end_year" is the last year of data we want to pull, so we loop to end_year+1
for year in range(start_year, end_year+1):
one_year_csv_writer(year, all_data)Because people will naturally expect that the end year for the files
is the last year with data, the for loop inside the function ends at
end_year + 1. By writing the entire loop into a function,
we’ve made a reusable tool for whenever we need to break a large data
file into yearly files. Because we can specify the first and last year
for which we want files, we can even use this function to create files
for a subset of the years available. This is how we call this
function:
PYTHON
# Load the data into a DataFrame
surveys_df = pd.read_csv('data/surveys.csv')
# Create CSV files
yearly_data_csv_writer(1977, 2002, surveys_df)BEWARE! If you are using Jupyter Notebooks and you modify a function, you MUST re-run that cell in order for the changed function to be available to the rest of the code. Nothing will visibly happen when you do this, though, because defining a function without calling it doesn’t produce an output. Any cells that use the now-changed functions will also have to be re-run for their output to change.
Challenge: More functions
- Add two arguments to the functions we wrote that take the path of the directory where the files will be written and the root of the file name. Create a new set of files with a different name in a different directory.
- How could you use the function
yearly_data_csv_writerto create a CSV file for only one year? (Hint: think about the syntax forrange) - Make the functions return a list of the files they have written.
There are many ways you can do this (and you should try them all!):
either of the functions can print to screen, either can use a return
statement to give back numbers or strings to their function call, or you
can use some combination of the two. You could also try using the
oslibrary to list the contents of directories. - Explore what happens when variables are declared inside each of the functions versus in the main (non-indented) body of your code. What is the scope of the variables (where are they visible)? What happens when they have the same name but are given different values?
PYTHON
def one_year_csv_writer(this_year, all_data, folder_to_save, root_name): """ Writes a csv file for data from a given year. Parameters --------- this_year : int year for which data is extracted all_data: pd.DataFrame DataFrame with multi-year data folder_to_save : str folder to save the data files root_name: str root of the filenames to save the data """ # Select data for the year surveys_year = all_data[all_data.year == this_year] # Write the new DataFrame to a csv file filename = os.path.join(folder_to_save, ''.join([root_name, str(this_year), '.csv'])) surveys_year.to_csv(filename) start_year = 1978 end_year = 2000 directory_name = 'different_directory' root_file_name = 'different_file_name' for year in range(start_year, end_year+1): one_year_csv_writer(year, surveys_df, directory_name, root_file_name)yearly_data_csv_writer(2000, 2000, surveys_df)-
Building a list and returning it:
PYTHON
def one_year_csv_writer(this_year, all_data): """ Writes a csv file for data from a given year. this_year -- year for which data is extracted all_data -- DataFrame with multi-year data """ # Select data for the year surveys_year = all_data[all_data.year == this_year] # Write the new DataFrame to a csv file filename = 'data/yearly_files/function_surveys' + str(this_year) + '.csv' surveys_year.to_csv(filename) return filename def yearly_data_csv_writer(start_year, end_year, all_data): """ Writes separate CSV files for each year of data. start_year -- the first year of data we want end_year -- the last year of data we want all_data -- DataFrame with multi-year data """ # "end_year" is the last year of data we want to pull, so we loop to end_year+1 output_files = [] for year in range(start_year, end_year+1): output_files.append(one_year_csv_writer(year, all_data)) return output_files print(yearly_data_csv_writer(2000, 2001, surveys_df))OUTPUT
['data/yearly_files/function_surveys2000.csv', 'data/yearly_files/function_surveys2001.csv']
The functions we wrote demand that we give them a value for every
argument. Ideally, we would like these functions to be as flexible and
independent as possible. Let’s modify the function
yearly_data_csv_writer so that the start_year
and end_year default to the full range of the data if they
are not supplied by the user. Arguments can be given default values with
an equal sign in the function declaration. Any arguments in the function
without default values (here, all_data) is a required
argument and MUST come before the argument with default values (which
are optional in the function call).
PYTHON
def yearly_data_arg_test(all_data, start_year=1977, end_year=2002):
"""
Modified from yearly_data_csv_writer to test default argument values!
start_year -- the first year of data we want (default 1977)
end_year -- the last year of data we want (default 2002)
all_data -- DataFrame with multi-year data
"""
return start_year, end_year
start, end = yearly_data_arg_test(surveys_df, 1988, 1993)
print('Both optional arguments:\t', start, end)
start, end = yearly_data_arg_test(surveys_df)
print('Default values:\t\t\t', start, end)OUTPUT
Both optional arguments: 1988 1993
Default values: 1977 2002The “\t” in the print statements are tabs, used to make
the text align and be easier to read.
But what if our dataset doesn’t start in 1977 and end in 2002? We can modify the function so that it looks for the start and end years in the dataset if those dates are not provided:
PYTHON
def yearly_data_arg_test(all_data, start_year=None, end_year=None):
"""
Modified from yearly_data_csv_writer to test default argument values!
all_data -- DataFrame with multi-year data
start_year -- the first year of data we want, Check all_data! (default None)
end_year -- the last year of data we want; Check all_data! (default None)
"""
if start_year is None:
start_year = min(all_data.year)
if end_year is None:
end_year = max(all_data.year)
return start_year, end_year
start, end = yearly_data_arg_test(surveys_df, 1988, 1993)
print('Both optional arguments:\t', start, end)
start, end = yearly_data_arg_test(surveys_df)
print('Default values:\t\t\t', start, end)OUTPUT
Both optional arguments: 1988 1993
Default values: 1977 2002The default values of the start_year and
end_year arguments in the function
yearly_data_arg_test are now None. This is a
built-in constant in Python that indicates the absence of a value -
essentially, that the variable exists in the namespace of the function
(the directory of variable names) but that it doesn’t correspond to any
existing object.
Challenge - Variables
- What type of object corresponds to a variable declared as
None? (Hint: create a variable set toNoneand use the functiontype()) - Compare the behavior of the function
yearly_data_arg_testwhen the arguments haveNoneas a default and when they do not have default values. - What happens if you only include a value for
start_yearin the function call? Can you write the function call with only a value forend_year? (Hint: think about how the function must be assigning values to each of the arguments - this is related to the need to put the arguments without default values before those with default values in the function definition!)
-
OUTPUT
NoneType -
Removing
Noneas the default value for thestart_yearandend_yearparameters results in aTypeErrorbecause the function now expects three arguments to be provided:PYTHON
def yearly_data_arg_test(all_data, start_year, end_year): """ Modified from yearly_data_csv_writer to test default argument values! all_data -- DataFrame with multi-year data start_year -- the first year of data we want, Check all_data! (default None) end_year -- the last year of data we want; Check all_data! (default None) """ if start_year is None: start_year = min(all_data.year) if end_year is None: end_year = max(all_data.year) return start_year, end_year start, end = yearly_data_arg_test(surveys_df)ERROR
TypeError: yearly_data_arg_test() missing 2 required positional arguments: 'start_year' and 'end_year' -
Returning to the original definition of the function (i.e. with the default
Nonevalues):OUTPUT
(1985, 2002)If we want to provide only the end year, we have to name the arguments:
OUTPUT
(1977, 1985)
If Statements
The body of the test function now has two conditionals (if
statements) that check the values of start_year and
end_year. If statements execute a segment of code when some
condition is met. They commonly look something like this:
PYTHON
a = 5
if a<0: # Meets first condition?
# if a IS less than zero
print('a is a negative number')
elif a>0: # Did not meet first condition. meets second condition?
# if a ISN'T less than zero and IS more than zero
print('a is a positive number')
else: # Met neither condition
# if a ISN'T less than zero and ISN'T more than zero
print('a must be zero!')Which would return:
OUTPUT
a is a positive numberChange the value of a to see how this function works.
The statement elif means “else if”, and all of the
conditional statements must end in a colon.
The if statements in the function yearly_data_arg_test
check whether there is an object associated with the variable names
start_year and end_year. If those variables
are None, the if statements return the boolean
True and execute whatever is in their body. On the other
hand, if the variable names are associated with some value (they got a
number in the function call), the if statements return
False and do not execute. The opposite conditional
statements, which would return True if the variables were
associated with objects (if they had received value in the function
call), would be if start_year and
if end_year.
As we’ve written it so far, the function
yearly_data_arg_test associates values in the function call
with arguments in the function definition just based on their order. If
the function gets only two values in the function call, the first one
will be associated with all_data and the second with
start_year, regardless of what we intended them to be. We
can get around this problem by calling the function using keyword
arguments, where each of the arguments in the function definition is
associated with a keyword and the function call passes values to the
function using these keywords:
PYTHON
start, end = yearly_data_arg_test(surveys_df)
print('Default values:\t\t\t', start, end)
start, end = yearly_data_arg_test(surveys_df, 1988, 1993)
print('No keywords:\t\t\t', start, end)
start, end = yearly_data_arg_test(surveys_df, start_year=1988, end_year=1993)
print('Both keywords, in order:\t', start, end)
start, end = yearly_data_arg_test(surveys_df, end_year=1993, start_year=1988)
print('Both keywords, flipped:\t\t', start, end)
start, end = yearly_data_arg_test(surveys_df, start_year=1988)
print('One keyword, default end:\t', start, end)
start, end = yearly_data_arg_test(surveys_df, end_year=1993)
print('One keyword, default start:\t', start, end)OUTPUT
Default values: 1977 2002
No keywords: 1988 1993
Both keywords, in order: 1988 1993
Both keywords, flipped: 1988 1993
One keyword, default end: 1988 2002
One keyword, default start: 1977 1993Challenge - Modifying functions
Rewrite the
one_year_csv_writerandyearly_data_csv_writerfunctions to have keyword arguments with default valuesModify the functions so that they don’t create yearly files if there is no data for a given year and display an alert to the user (Hint: use conditional statements to do this. For an extra challenge, use
trystatements!)-
The code below checks to see whether a directory exists and creates one if it doesn’t. Add some code to your function that writes out the CSV files, to check for a directory to write to.
The code that you have written so far to loop through the years is good, however it is not necessarily reproducible with different datasets. For instance, what happens to the code if we have additional years of data in our CSV files? Using the tools that you learned in the previous activities, make a list of all years represented in the data. Then create a loop to process your data, that begins at the earliest year and ends at the latest year using that list.
HINT: you can create a loop with a list as follows:
for years in year_list:
PYTHON
def one_year_csv_writer(this_year, all_data, folder_to_save='./', root_name='survey'): """ Writes a csv file for data from a given year. Parameters --------- this_year : int year for which data is extracted all_data: pd.DataFrame DataFrame with multi-year data folder_to_save : str folder to save the data files root_name: str root of the filenames to save the data """ # Select data for the year surveys_year = all_data[all_data.year == this_year] # Write the new DataFrame to a csv file filename = os.path.join(folder_to_save, ''.join([root_name, str(this_year), '.csv'])) surveys_year.to_csv(filename) def yearly_data_csv_writer(all_data, start_year=None, end_year=None): """ Writes separate CSV files for each year of data. start_year -- the first year of data we want end_year -- the last year of data we want all_data -- DataFrame with multi-year data """ if start_year is None: start_year = min(all_data.year) if end_year is None: end_year = max(all_data.year) # "end_year" is the last year of data we want to pull, so we loop to end_year+1 for year in range(start_year, end_year+1): one_year_csv_writer(year, all_data)PYTHON
def one_year_csv_writer(this_year, all_data, folder_to_save='./', root_name='survey'): """ Writes a csv file for data from a given year. Parameters --------- this_year : int year for which data is extracted all_data: pd.DataFrame DataFrame with multi-year data folder_to_save : str folder to save the data files root_name: str root of the filenames to save the data """ # Select data for the year surveys_year = all_data[all_data.year == this_year] if len(surveys_year) == 0: # 'if not len(surveys_year):' will also work print('no data available for ' + this_year + ', output file not created') else: # Write the new DataFrame to a csv file filename = os.path.join(folder_to_save, ''.join([root_name, str(this_year), '.csv'])) surveys_year.to_csv(filename)-
The solution below uses
notto invert the check for the directory, so a message is printed one time at most i.e. when the directory is created:PYTHON
def one_year_csv_writer(this_year, all_data, folder_to_save='./', root_name='survey'): """ Writes a csv file for data from a given year. Parameters --------- this_year : int year for which data is extracted all_data: pd.DataFrame DataFrame with multi-year data folder_to_save : str folder to save the data files root_name: str root of the filenames to save the data """ # Select data for the year surveys_year = all_data[all_data.year == this_year] if len(surveys_year) == 0: print('no data available for ' + this_year + ', output file not created') else: if folder_to_save not in os.listdir('.'): os.mkdir(folder_to_save) print('Processed directory created') # Write the new DataFrame to a csv file filename = os.path.join(folder_to_save, ''.join([root_name, str(this_year), '.csv'])) surveys_year.to_csv(filename)The equivalent, without using
notto invert the conditional, would be: PYTHON
def yearly_data_csv_writer(all_data, yearcolumn="year", folder_to_save='./', root_name='survey'): """ Writes separate csv files for each year of data. all_data --- DataFrame with multi-year data yearcolumn --- column name containing the year of the data folder_to_save --- folder name to store files root_name --- start of the file names stored """ years = all_data[yearcolumn].unique() filenames = [] for year in years: filenames.append(one_year_csv_writer(year, all_data, folder_to_save, root_name)) return filenames
Key Points
- Loops help automate repetitive tasks over sets of items.
- Loops combined with functions provide a way to process data more efficiently than we could by hand.
- Conditional statements enable execution of different operations on different data.
- Functions enable code reuse.
Content from Making Plots With plotnine
Last updated on 2023-05-19 | Edit this page
Estimated time: 90 minutes
Overview
Questions
- How can I visualize data in Python?
- What is ‘grammar of graphics’?
Objectives
- Create a
plotnineobject. - Set universal plot settings.
- Modify an existing plotnine object.
- Change the aesthetics of a plot such as color.
- Edit the axis labels.
- Build complex plots using a step-by-step approach.
- Create scatter plots, box plots, and time series plots.
- Use the facet_wrap and facet_grid commands to create a collection of plots splitting the data by a factor variable.
- Create customized plot styles to meet their needs.
Disclaimer
Python has powerful built-in plotting capabilities such as
matplotlib, but for this episode, we will be using the plotnine package,
which facilitates the creation of highly-informative plots of structured
data based on the R implementation of ggplot2 and The
Grammar of Graphics by Leland Wilkinson. The plotnine
package is built on top of Matplotlib and interacts well with
Pandas.
Reminder
plotnine is not included in the standard Anaconda
installation and needs to be installed separately. If you haven’t done
so already, you can find installation instructions on the Setup
page.
Just as with the other packages, plotnine needs to be
imported. It is good practice to not just load an entire package such as
from plotnine import *, but to use an abbreviation as we
used pd for Pandas:
From now on, the functions of plotnine are available
using p9.. For the exercise, we will use the
surveys.csv data set, with the NA values
removed
PYTHON
import pandas as pd
surveys_complete = pd.read_csv('data/surveys.csv')
surveys_complete = surveys_complete.dropna()If learners have trouble generating the output, or anything happens
with that, the folder sample_output
in this repository contains the file surveys_complete.csv
with the data they should generate.
Plotting with plotnine
The plotnine package (cfr. other packages conform The
Grammar of Graphics) supports the creation of complex plots from data in
a dataframe. It uses default settings, which help creating publication
quality plots with a minimal amount of settings and tweaking.
plotnine graphics are built step by step by adding new
elements adding different elements on top of each other using the
+ operator. Putting the individual steps together in
brackets () provides Python-compatible syntax.
To build a plotnine graphic we need to:
- Bind the plot to a specific data frame using the
dataargument:
As we have not defined anything else, just an empty figure is available and presented.
- Define aesthetics (
aes), by selecting variables used in the plot andmappingthem to a presentation such as plotting size, shape, color, etc. You can interpret this as: which of the variables will influence the plotted objects/geometries:
The most important aes mappings are: x, y,
alpha, color, colour,
fill, linetype, shape,
size and stroke.
- Still no specific data is plotted, as we have to define what kind of
geometry will be used for the plot. The most straightforward is probably
using points. Points is one of the
geomsoptions, the graphical representation of the data in the plot. Others are lines, bars,… To add a geom to the plot use+operator:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length'))
+ p9.geom_point()
)The + in the plotnine package is
particularly useful because it allows you to modify existing
plotnine objects. This means you can easily set up plot
templates and conveniently explore different types of plots, so
the above plot can also be generated with code like this:
PYTHON
# Create
surveys_plot = p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length'))
# Draw the plot
surveys_plot + p9.geom_point()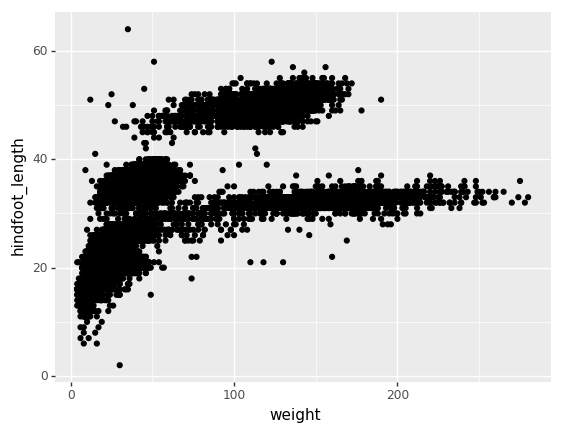
Challenge - bar chart
Working on the surveys_complete data set, use the
plot-id column to create a bar-plot that
counts the number of records for each plot. (Check the documentation of
the bar geometry to handle the counts)
Notes:
- Anything you put in the
ggplot()function can be seen by any geom layers that you add (i.e., these are universal plot settings). This includes thexandyaxis you set up inaes(). - You can also specify aesthetics for a given
geomindependently of the aesthetics defined globally in theggplot()function.
Building your plots iteratively
Building plots with plotnine is typically an iterative
process. We start by defining the dataset we’ll use, lay the axes, and
choose a geom. Hence, the data, aes and
geom-* are the elementary elements of any graph:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length'))
+ p9.geom_point()
)Then, we start modifying this plot to extract more information from it. For instance, we can add transparency (alpha) to avoid overplotting:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length'))
+ p9.geom_point(alpha=0.1)
)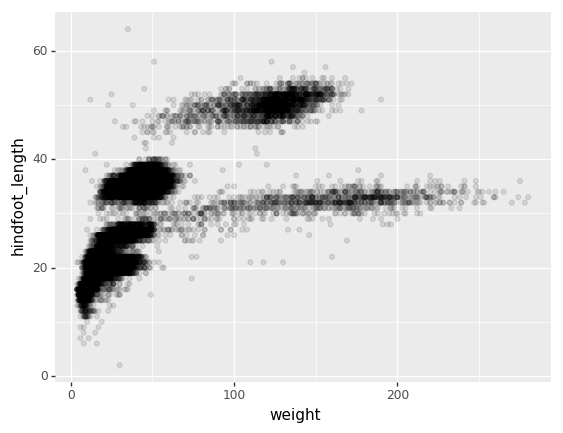
We can also add colors for all the points
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length'))
+ p9.geom_point(alpha=0.1, color='blue')
)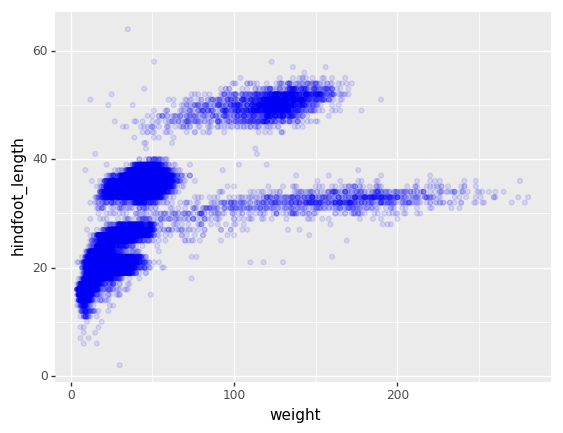
Or to color each species in the plot differently, map the
species_id column to the color aesthetic:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight',
y='hindfoot_length',
color='species_id'))
+ p9.geom_point(alpha=0.1)
)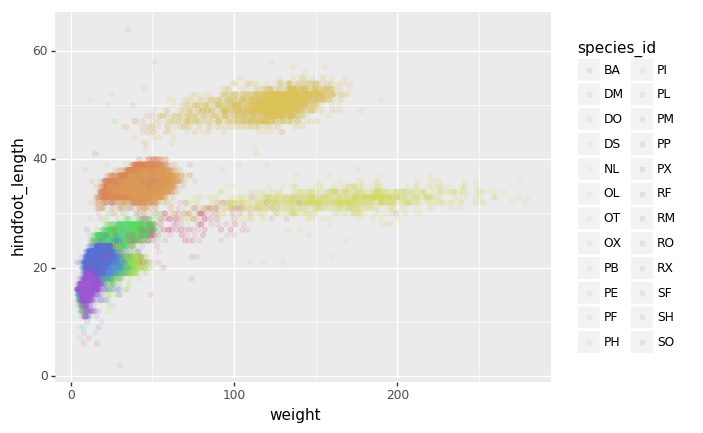
Apart from the adaptations of the arguments and settings of the
data, aes and geom-* elements,
additional elements can be added as well, using the +
operator:
- Changing the labels:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length', color='species_id'))
+ p9.geom_point(alpha=0.1)
+ p9.xlab("Weight (g)")
)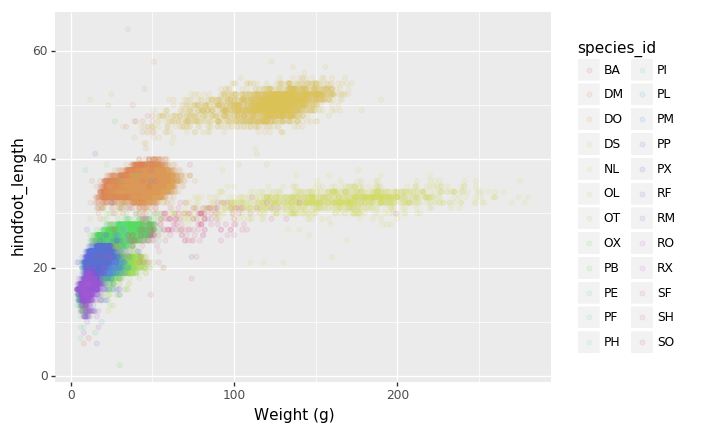
- Defining scale for colors, axes,… For example, a log-version of the x-axis could support the interpretation of the lower numbers:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length', color='species_id'))
+ p9.geom_point(alpha=0.1)
+ p9.xlab("Weight (g)")
+ p9.scale_x_log10()
)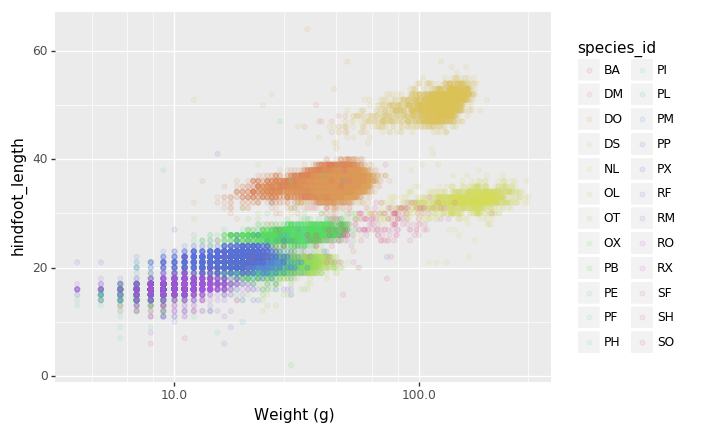
- Changing the theme (
theme_*) or some specific theming (theme) elements. Usually plots with white background look more readable when printed. We can set the background to white using the functiontheme_bw().
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length', color='species_id'))
+ p9.geom_point(alpha=0.1)
+ p9.xlab("Weight (g)")
+ p9.scale_x_log10()
+ p9.theme_bw()
+ p9.theme(text=p9.element_text(size=16))
)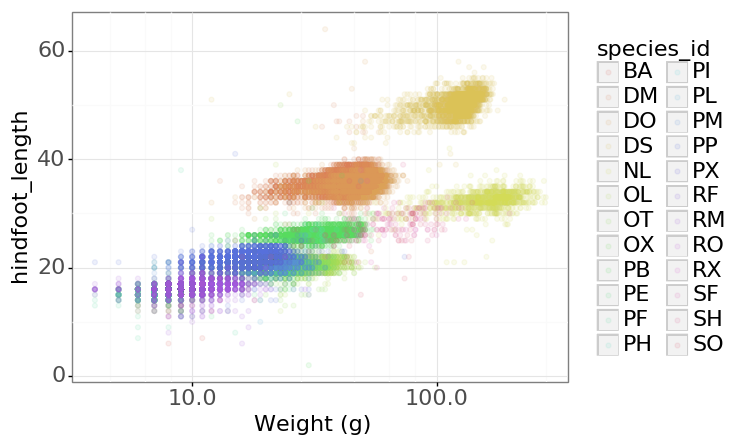
Challenge - Bar plot adaptations
Adapt the bar plot of the previous exercise by mapping the
sex variable to the color fill of the bar chart. Change the
scale of the color fill by providing the colors
blue and orange manually (see API
reference to find the appropriate function).
Plotting distributions
Visualizing distributions is a common task during data exploration
and analysis. To visualize the distribution of weight
within each species_id group, a boxplot can be used:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='species_id',
y='weight'))
+ p9.geom_boxplot()
)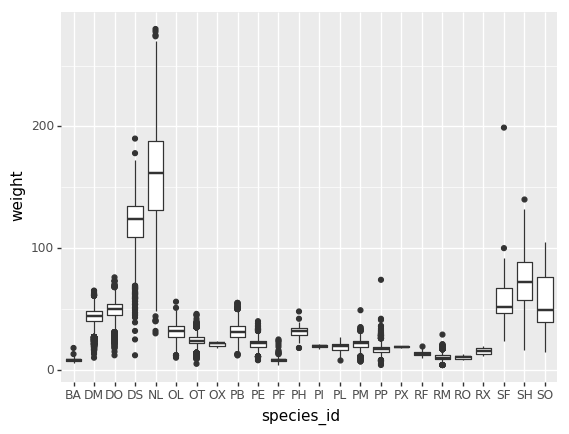
By adding points of the individual observations to the boxplot, we can have a better idea of the number of measurements and of their distribution:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='species_id',
y='weight'))
+ p9.geom_jitter(alpha=0.2)
+ p9.geom_boxplot(alpha=0.)
)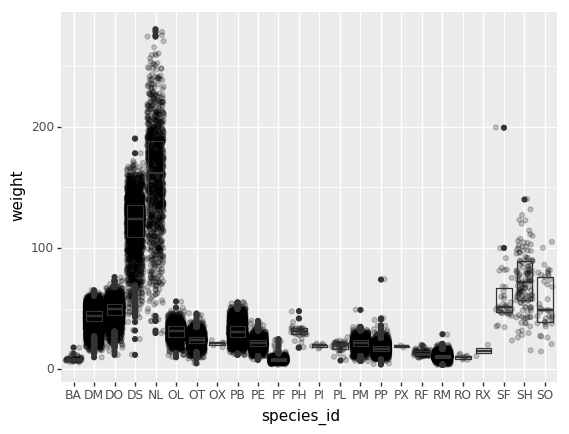
Challenge - distributions
Boxplots are useful summaries, but hide the shape of the distribution. For example, if there is a bimodal distribution, this would not be observed with a boxplot. An alternative to the boxplot is the violin plot (sometimes known as a beanplot), where the shape (of the density of points) is drawn.
In many types of data, it is important to consider the scale of the observations. For example, it may be worth changing the scale of the axis to better distribute the observations in the space of the plot.
- Replace the box plot with a violin plot, see
geom_violin() - Represent weight on the log10 scale, see
scale_y_log10() - Add color to the datapoints on your boxplot according to the plot
from which the sample was taken (
plot_id)
Hint: Check the class for plot_id. By using
factor() within the aes mapping of a variable,
plotnine will handle the values as category values.
Plotting time series data
Let’s calculate number of counts per year for each species. To do
that we need to group data first and count the species
(species_id) within each group.
PYTHON
yearly_counts = surveys_complete.groupby(['year', 'species_id'])['species_id'].count()
yearly_countsWhen checking the result of the previous calculation, we actually
have both the year and the species_id as a row
index. We can reset this index to use both as column variable:
Timelapse data can be visualised as a line plot
(geom_line) with years on x axis and counts on
the y axis.
Unfortunately this does not work, because we plot data for all the
species together. We need to tell plotnine to draw a line
for each species by modifying the aesthetic function and map the
species_id to the color:
PYTHON
(p9.ggplot(data=yearly_counts,
mapping=p9.aes(x='year',
y='counts',
color='species_id'))
+ p9.geom_line()
)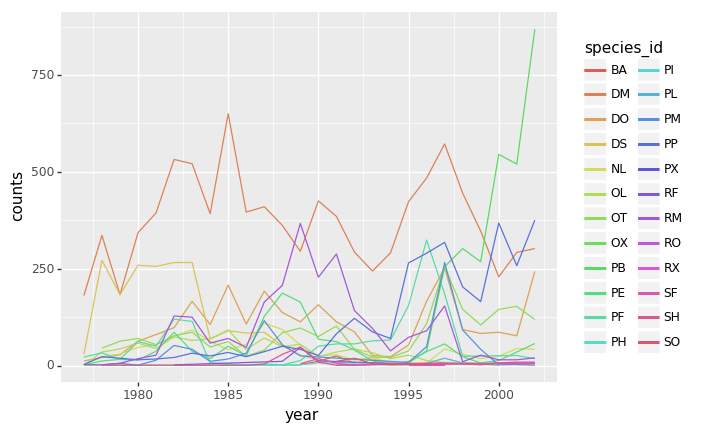
Faceting
As any other library supporting the Grammar of Graphics,
plotnine has a special technique called faceting
that allows to split one plot into multiple plots based on a factor
variable included in the dataset.
Consider our scatter plot of the weight versus the
hindfoot_length from the previous sections:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight',
y='hindfoot_length',
color='species_id'))
+ p9.geom_point(alpha=0.1)
)We can now keep the same code and at the facet_wrap on a
chosen variable to split out the graph and make a separate graph for
each of the groups in that variable. As an example, use
sex:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight',
y='hindfoot_length',
color='species_id'))
+ p9.geom_point(alpha=0.1)
+ p9.facet_wrap("sex")
)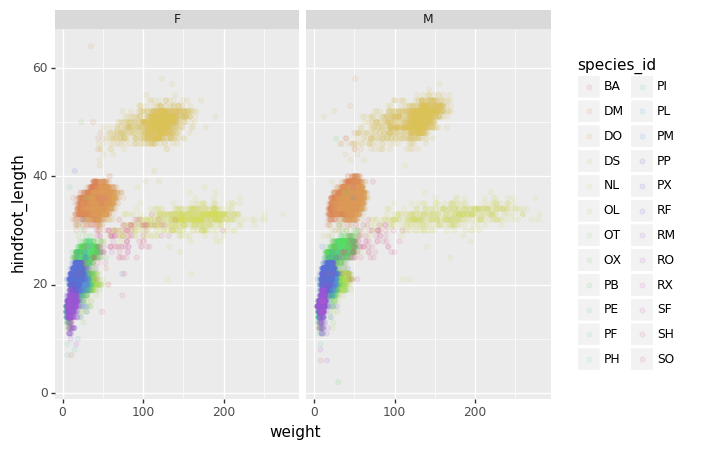
We can apply the same concept on any of the available categorical variables:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight',
y='hindfoot_length',
color='species_id'))
+ p9.geom_point(alpha=0.1)
+ p9.facet_wrap("plot_id")
)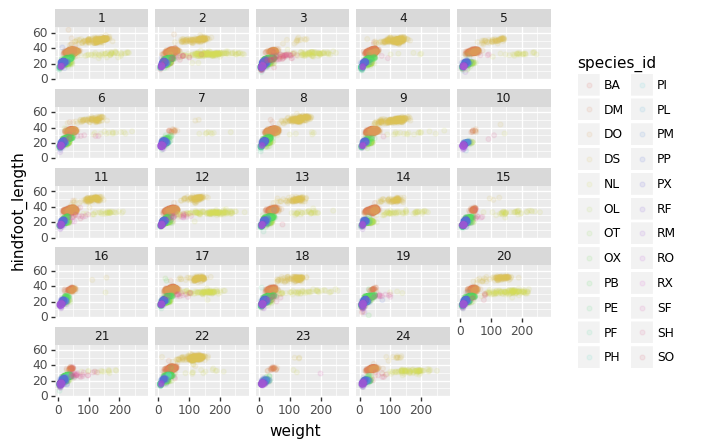
The facet_wrap geometry extracts plots into an arbitrary
number of dimensions to allow them to cleanly fit on one page. On the
other hand, the facet_grid geometry allows you to
explicitly specify how you want your plots to be arranged via formula
notation (rows ~ columns; a . can be used as a
placeholder that indicates only one row or column).
PYTHON
# only select the years of interest
survey_2000 = surveys_complete[surveys_complete["year"].isin([2000, 2001])]
(p9.ggplot(data=survey_2000,
mapping=p9.aes(x='weight',
y='hindfoot_length',
color='species_id'))
+ p9.geom_point(alpha=0.1)
+ p9.facet_grid("year ~ sex")
)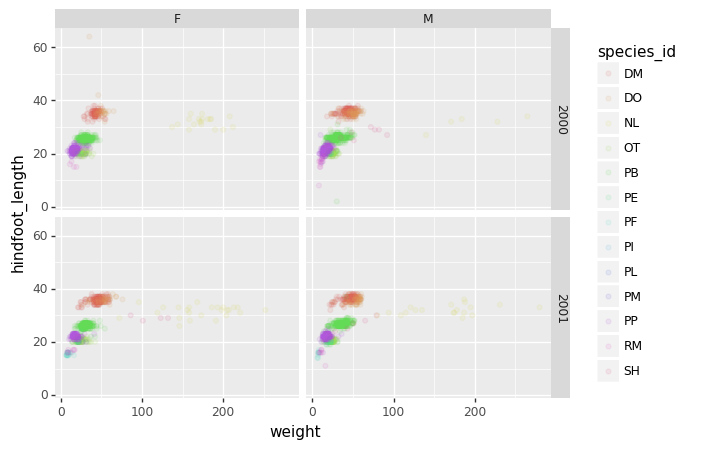
Challenge - facetting
Create a separate plot for each of the species that depicts how the average weight of the species changes through the years.
Challenge - facetting
Based on the previous exercise, visually compare how the weights of
male and females has changed through time by creating a separate plot
for each sex and an individual color assigned to each
species_id.
Further customization
As the syntax of plotnine follows the original R package
ggplot2, the documentation of ggplot2 can
provide information and inspiration to customize graphs. Take a look at
the ggplot2 cheat
sheet, and think of ways to improve the plot. You can write down
some of your ideas as comments in the Etherpad.
The theming options provide a rich set of visual adaptations. Consider the following example of a bar plot with the counts per year.
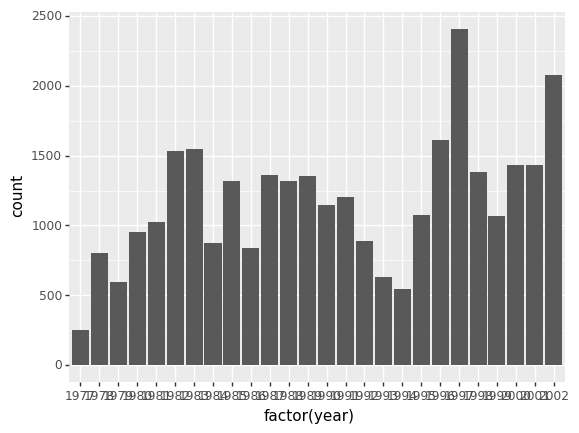
Notice that we use the year here as a categorical
variable by using the factor functionality. However, by
doing so, we have the individual year labels overlapping with each
other. The theme functionality provides a way to rotate the
text of the x-axis labels:
PYTHON
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='factor(year)'))
+ p9.geom_bar()
+ p9.theme_bw()
+ p9.theme(axis_text_x = p9.element_text(angle=90))
)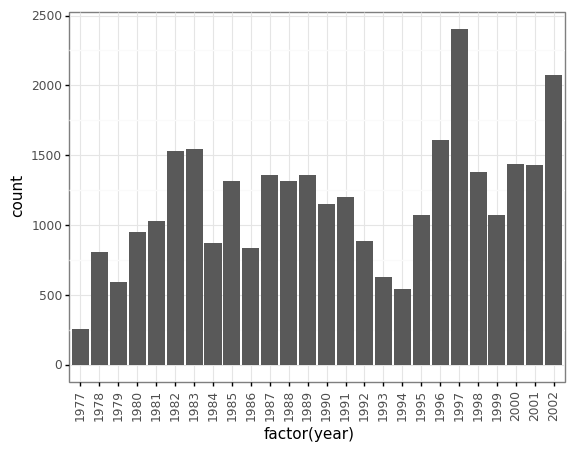
When you like a specific set of theme-customizations you created, you can save them as an object to easily apply them to other plots you may create:
PYTHON
my_custom_theme = p9.theme(axis_text_x = p9.element_text(color="grey", size=10,
angle=90, hjust=.5),
axis_text_y = p9.element_text(color="grey", size=10))
(p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='factor(year)'))
+ p9.geom_bar()
+ my_custom_theme
)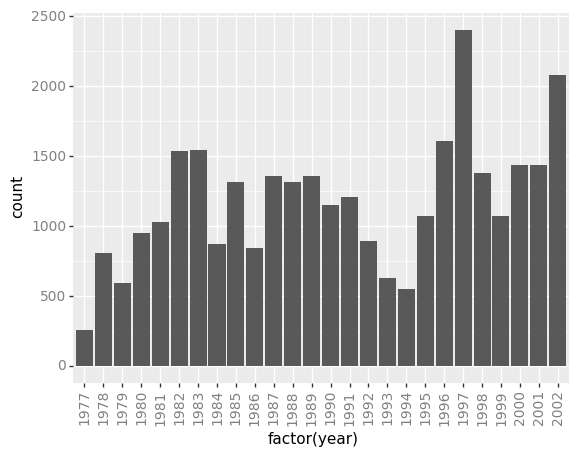
Challenge - customization
Please take another five minutes to either improve one of the plots generated in this exercise or create a beautiful graph of your own.
Here are some ideas:
- See if you can change thickness of lines for the line plot .
- Can you find a way to change the name of the legend? What about its labels?
- Use a different color palette (see http://www.cookbook-r.com/Graphs/Colors_(ggplot2))
After creating your plot, you can save it to a file in your favourite
format. You can easily change the dimension (and its resolution) of your
plot by adjusting the appropriate arguments (width,
height and dpi):
PYTHON
my_plot = (p9.ggplot(data=surveys_complete,
mapping=p9.aes(x='weight', y='hindfoot_length'))
+ p9.geom_point()
)
my_plot.save("scatterplot.png", width=10, height=10, dpi=300)Key Points
- The
data,aesvariables and ageometryare the main elements of a plotnine graph - With the
+operator, additionalscale_*,theme_*,xlab/ylabandfacet_*elements are added